Products
In the context of the GeneReader NGS System, the QIAsymphony SP, QIAsymphony DPS DNA Mini Kit and QIAsymphony GeneRead FFPE Treatment Kits are intended for Research Use Only. Not for use in diagnostic procedures.

The QIAsymphony DSP DNA Mini Kit (192) is intended for in vitro diagnostic use.
Features
- Rapid purification of high-quality, ready-to-use DNA for use in research applications
- Enzymatic removal of cytosine deamination artifacts for FFPE samples
- Automation on the QIAsymphony Sample Preparation
- Outstanding results in DNA sequencing applications
Product Details
With the QIAsymphony DSP DNA Mini Kit, in combination with the QIAsymphony GeneRead DNA FFPE Treatment Kit and QIAsymphony SP, it is possible to fully automate DNA extraction from up to 96 FFPE samples in parallel. Enzymatic removal of cytosine deamination artifacts from FFPE samples in combination with optimized protocols provide high yields of pure DNA, suitable for downstream sequencing applications in the GeneReader NGS System workflow. Novel, prefilled reagent cartridges reduce setup time and minimize manual handling. Bar code reading enables full tracking of reagents.
Principle
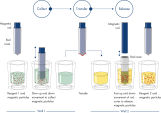
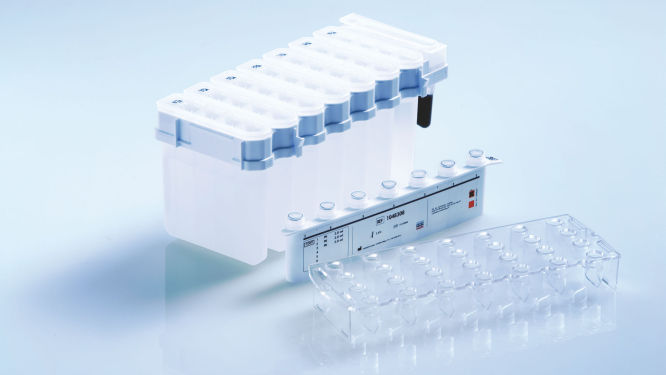
The QIAsymphony DSP DNA mini kit in combination with the QIAsymphony GeneRead DNA FFPE Treatment Kit can be used to purify DNA from FFPE tissues. This combines the speed and efficiency of silica-based purification of DNA with the convenient handling of magnetic particles (see figure " Schematic of the QIAsymphony SP principle"). Innovative, ready-to-run reagent cartridges are prefilled with all reagents required for the purification procedure and are automatically opened by the QIAsymphony SP, which helps to minimize the risk of environmental contamination (see figure " Prefilled, sealed reagent cartridges"). Worktable setup is rapid, which saves you valuable time. Simply place up to 2 reagent cartridges in the consumables drawer. The reagent cartridges can be either from the same kit or from different kits, enabling different purification procedures to be performed within the same run of 96 samples. Reagent volumes only for the selected number of samples are used, giving you complete cost control.
Procedure
The purification procedure is designed to ensure safe and reproducible handling of potentially infectious samples, and comprises 4 steps: lyse, bind, wash, and elute (see figure " QIAsymphony DSP DNA procedure"). Optimized protocols provide high quality nucleic acids with efficient removal of proteins, nucleases, and other impurities. Nucleic acids purified with QIAsymphony GeneRead DNA FFPE Treatment Kit and QIAsymphony DSP DNA Mini Kit is ready for direct use in downstream library preparation and sequencing applications on the GeneReader NGS System. The user can choose between different elution volumes.
Applications
Designed for use in the GeneReader NGS System workflow.
Supporting data and figures
Prefilled, sealed reagent cartridges.
.
Service Plans
Resources
Kit Handbooks (2)
Brochures and Guides (6)
Labware File Documents (3)
Protocols (47)
Scientific Posters (4)
Supporting Software and Tools (2)
Technical Information (1)
User Manuals (15)
Validation Reports (2)
Performance Data (1)
Safety Data Sheets (1)
Certificates of Analysis (1)