QIAexpress Type IV Kit
N末端Hisタグ付加タンパク質の高レベル発現およびワンステップ精製のために
N末端Hisタグ付加タンパク質の高レベル発現およびワンステップ精製のために
✓ オンライン注文による24時間年中無休の自動処理システム
✓ 知識豊富で専門的な製品&テクニカルサポート
✓ 迅速で信頼性の高い(再)注文
Cat. No. / ID: 32149
✓ オンライン注文による24時間年中無休の自動処理システム
✓ 知識豊富で専門的な製品&テクニカルサポート
✓ 迅速で信頼性の高い(再)注文
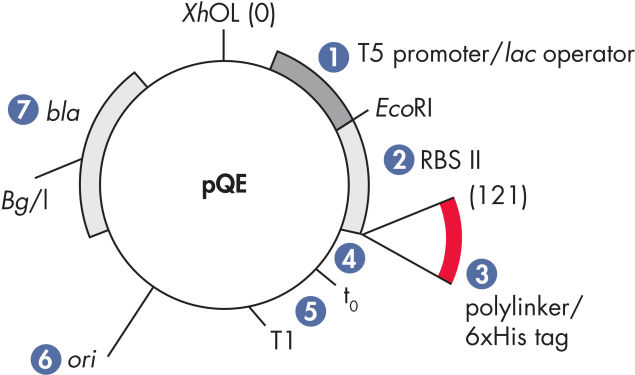
QIAexpress pQEベクターは、強力なファージT5プロモーター(E. coli RNAポリメラーゼによって認識)を二重のlacオペレーター抑制モジュールと組み合わせて、E. coliで遺伝子組み換えタンパク質の厳しく制御される、高レベルな発現を実現します。タンパク質合成は、高レベルのlacリプレッサーの存在で効果的にブロックされ、細胞毒性コンストラクトの安定性が向上します。pQEベクター( 図 pQE Vectors および表を参照)は、遺伝子組み換えタンパク質のN末端またはC末端のいずれかの Hisタグの配置を可能にします。
| 要素 | 説明 |
|---|---|
| 1. 最適化されたプロモーター/オペレーターエレメント | ファージT5プロモーターと2つのlacオペレーター配列で構成され、それによりlacリプレッサー結合の可能性が上がり、強力なT5プロモーターの効率的な抑制を確実にします |
| 2. 合成リボソーム結合部位RBSII | 効率的な翻訳のために |
| 3. Hisタグをコードする配列 | ポリリンカークローニング領域への5’または3’のいずれか |
| 4. 翻訳終止コドン | 発現コンストラクトの便利な調製のため、すべてのリーディングフレーム内 |
| 5. 2つの強力な転写ターミネーター | λファージ由来のt0、およびE. coliのrrnBオペロン由来のT1はリードする―転写を防ぎ、発現コンストラクトの安定性を確保する |
| 6. ColE1複製開始点 | pBR322由来 |
| 7. βラクタマーゼ遺伝子 (bla) | アンピシリン耐性を与えます |
目的のタンパク質をコードするインサートは、適切なコンストラクト内にクローンされ、発現のために適切なE. coli株内に形質転換されます。発現は、IPTGの添加によって誘導されます。QIAexpress Type IV Kitのコンストラクトは、E.coli内に転換されたり、昆虫細胞内での遺伝子組み換えタンパク質の発現のためにシャトルベクターとして使用されたり、哺乳類細胞内に導入されたりします。
QIAexpress Expression Systemは、以下をはじめとする数多くのアプリケーションに適したタンパク質の高レベル発現を実現します。
| Features | Specifications |
|---|---|
| Applications | プロテオミクス |
| Yield | 総タンパク質の50%未満 |
| 特殊機能 | ワンステップ精製のための多用途で完全なシステム |
| Tag | 6xHisタグ |
| Processing | 手動 |
| Start material | 細胞溶解物 |
| N- or C-terminal tag | N末端タグ |