Configure at GeneGlobe
Find or custom design the right target-specific assays and panels to research your biological targets of interest.
miRCURY LNA miRNA Power Family Inhibitors
Cat. No. / ID: 339160
Predesigned sets of miRCURY LNA miRNA Inhibitors for miRNA families
Configure at GeneGlobe To see pricing
miRCURY LNA miRNA Family Power Inhibitors are intended for molecular biology applications. These products are not intended for the diagnosis, prevention, or treatment of a disease.
Configure at GeneGlobe
Find or custom design the right target-specific assays and panels to research your biological targets of interest.
Features
- Enable functional analysis of an entire miRNA family, instead of just individual family members
- LNA design ensures that all miRNA family members are silenced with similar high efficacy, regardless of GC content
- More than 40 predesigned inhibitors of miRNA families available
- Allows discovery of novel miRNA functions that would not be revealed when analyzing individual family members
Product Details
miRCURY LNA miRNA Family Power Inhibitors allow you to study regulatory roles shared by highly related, co-expressed and functionally redundant miRNAs. Such functions would not be revealed in analyses using inhibitors of individual family members. These antisense oligonucleotides have perfect sequence complementarity to their targets, and when introduced into cells, they sequester the target miRNA in highly stable heteroduplexes that prevent interaction with the normal cellular partners.
Need a quote for your research project or would you like to discuss your project with our specialist team? Just contact us!
Performance
Discover unknown miRNA function
miRNA family members are often co-expressed, either from the same pri-miR transcript or from distinct but co-regulated loci. This suggests that miRNAs within a family may share a common regulatory function. Given their identical seed sequence, miRNA family members will have many mRNA targets in common, and they are therefore likely to be functionally redundant. Therefore, analysis of miRNA family function requires simultaneous inhibition of all or several family members.The miRCURY LNA miRNA Family Power Inhibitors allow you to observe robust phenotypes that would be either absent or weakly presented if using inhibitors of individual miRNA family members (see figure The miR-30 family positively regulates hMADS cell adipocyte differentiation). This enables you to discover new, unknown miRNA functions that you may have deduced through careful analysis of miRNA target predictions, but have not been able to prove experimentally.
Principle
Design
miRCURY LNA miRNA Family Power Inhibitors are specifically designed to silence entire families of miRNAs, rather than individual family members. Through careful design, we have developed special family inhibitors that knock down all family members, while retaining specificity for the targeted family (see figure Example of miRNA family inhibitor design). This is achieved through the following design criteria:- Sequence optimization: identification of a target sequence sufficiently short to be shared by family members and sufficiently long to define the family.
- Mixed-base synthesis: when target sequences differ at a single position, we use a mixture of two bases at the corresponding position during the synthesis of the antisense oligonucleotide. This allows us to generate two oligonucleotides in a single oligonucleotide synthesis reaction.
- Mixture of inhibitors: when it is not possible to target a whole family using a single nucleotide synthesis, we provide a mixture of two or more inhibitors that make up the family inhibitor.
- LNA optimization: family inhibitors are short 10–16mers, but enhanced LNA content ensures that they all have the same target affinity (Tm) as our individual miRCURY LNA miRNA inhibitors.
Coverage
We have developed inhibitors for more than 40 human miRNA families that are conserved in mouse. miRCURY LNA miRNA Family Power Inhibitors have been designed only for miRNA families in which the design criteria described above provide an advantage over simply using a mixture of individual inhibitors of all miRNA family members.We define miRNA families as highly related miRNAs that share an identical seed sequence (nucleotides 2–8). This is particularly important for target recognition, although the miR-200 family is one of the few exceptions to this rule.
Some very large, miRBase-defined miRNA families, such as the hsa-miR-506, hsa-miR-515 and hsa-miR-548 families, comprise miRNAs with similar sequences but with different seed sequences. These families are not represented in our miRCURY LNA miRNA Family Power Inhibitors.
Power Inhibitors are not recommended for use with cells or cell lines derived from muscle or the central nervous system (CNS). These cell types are known to be particularly sensitive to sequence-dependent toxicity of phosphorothioate-modified oligonucleotides.
See figures
Procedure
miRCURY LNA miRNA Family Power Inhibitors are antisense oligonucleotides with perfect sequence complementarity to their targets. When introduced into cells, they sequester the target miRNA in highly stable heteroduplexes, effectively preventing the miRNA from hybridizing with its normal cellular interaction partners. The sequences of the oligonucleotides and their LNA spiking patterns have been carefully designed to achieve uniform high potency, regardless of the GC content of the target. The Tm is normalized to an optimal temperature, and the level of self-complementarity is kept to a minimum.
Following resuspension, miCURY LNA miRNA Family Power Inhibitors are introduced into cells with a transfection reagent or via electroporation. Phenotypic effects are at an appropriate time thereafter.
Following resuspension, miCURY LNA miRNA Family Power Inhibitors are introduced into cells with a transfection reagent or via electroporation. Phenotypic effects are at an appropriate time thereafter.
Applications
miRCURY LNA miRNA Inhibitors are primarily used miRNA functional studies by assessing the biological consequences of inhibiting miRNA activity. These effects can be assessed in a variety of ways, including using cellular assays to monitor cell proliferation, cell differentiation or apoptosis. The effects on gene expression can also be measured at the mRNA or protein level of putative miRNA targets.
Supporting data and figures
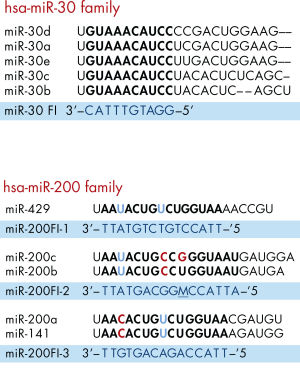
Example of miRNA family inhibitor design.
Example of miRNA family inhibitor design. (A) miR-30 family: identification of a shared sequence unique for the miR-30 family (in bold). In this particular case it was possible to design one short, complementary inhibitor, miR-30 Family Inhibitor (FI), with high duplex melting temperature (Tm) that binds with high specificity to members of the miR-30 miRNA family. (B) miR-200 family: silencing of the miR-200 family cannot be achieved with a single inhibitor. However, the five family members can be addressed with three inhibitors. The miR-200 Family Inhibitor (FI)-2 is the result of an oligonucleotide synthesis with a mixed-base composition, giving rise to two oligonucleotides that differ at position 7 (from the 5’ end). M = A,C. Letters in color indicate positions in which the target sequence varies between family members.
Resources
Safety Data Sheets (1)
Kit Handbooks (1)
Brochures & Guides (1)
Certificates of Analysis (1)



