Omniscript RT Kit
For reverse transcription of 50 ng to 2 μg RNA per reaction for end-point PCR
For reverse transcription of 50 ng to 2 μg RNA per reaction for end-point PCR
Cat. No. / ID: 205113
The high affinity of Omniscript Reverse Transcriptase (RT) results in highly specific and sensitive RT-PCR, even with low-copy transcripts (see figure " Sensitive RT-PCR of ≥10 copies"), and the ability to read through even complex RNA secondary structures without adjusting temperature or reaction conditions (see figure " Comparison of various reverse transcriptases").
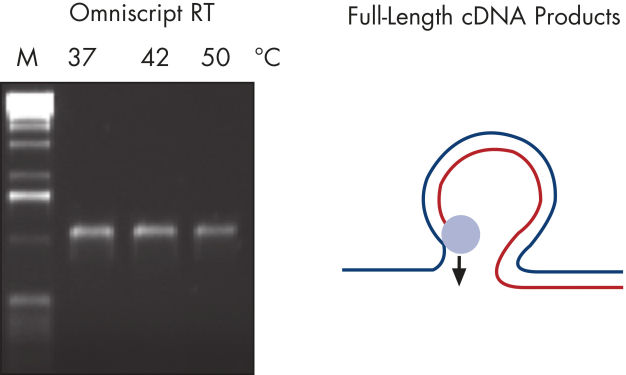
Regions of RNA with high GC content can cause other reverse transcriptases to stop, dissociate from the RNA template, or skip over looped-out regions of RNA (see figure " Full-length RT-PCR products — B"). These difficult templates, however, prove no problem for QIAGEN reverse transcriptases (see figure " Full-length RT-PCR products — A"). With no need for optimization, the Omniscript RT Kit makes reverse transcription virtually trouble-free.
Omniscript Reverse Transcriptase (RT) has a high affinity for RNA, which enables efficient and sensitive reverse transcription of any template, leading to high yields of cDNA. It is provided ready-to-use with dNTPs and in an optimized reaction buffer that, together with the high affinity of Omniscript RT for RNA, enables read-through of templates with high GC content or complex secondary structure. Please note that the primer mix is not provided.
Omniscript RT is specially designed for all reverse transcription with any amount of RNA from 50 ng to 2 µg per reaction. Omniscript RT is also usually the enzyme of choice with viral RNA due to the presence of carrier RNA in most viral RNA preparations. In comparative experiments, Omniscript RT consistently outperforms other reverse transcriptases over a wide range of starting RNA amounts.
Lot-to-lot reproducibility of Omniscript Kits is ensured by rigorous quality control at QIAGEN. The optimized Buffer RT, dNTPs, and water included in all Omniscript RT Kits are guaranteed RNase-free, and each lot of Omniscript RT is thoroughly tested for RT-PCR reproducibility.
Omniscript Reverse Transcriptase is suitable for use in the following applications:
| Features | Specifications |
|---|---|
| Applications | RT-PCR, qRT-PCR, primer-extension, RACE analysis |
| Real-time or endpoint | Endpoint |
| Mastermix | No |
| Enzyme activity | Reverse transcription |
| Single or multiplex | Single |
| With/without hotstart | Without hotstart |
| Reaction type | Reverse transcription |
| Sample/target type | RNA template |