✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Cat. No. / ID: 32903
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- La etiqueta 6xHis del extremo C-terminal garantiza que solo se purifica la proteína de duración completa
- Vector pQE-16 para la expresión de proteínas poco expresadas como fusiones de DHFR
- Vector pQE-16 para la expresión de péptidos cortos como fusiones de DHFR
Product Details
Este conjunto proporciona 3 vectores (pQE-16, pQE-60 y pQE-70) para la expresión de proteínas de extremo C-terminal marcadas con 6xHis y está recomendado para los marcos de lectura abiertos con «sitios de pausa», que pueden causar una finalización prematura. pQE-60 y pQE-70 permiten que el codón original de inicio del fragmento de codificación sustituya al ATG en el vector pQE, conservando el N-terminal auténtico de la proteína. Estas dos construcciones se crean introduciendo un sitio de restricción NcoI y SphI, respectivamente, en el codón ATG del inserto mediante PCR o mutagénesis. pQE-16 permite la expresión de proteínas de fusión de DHFR de extremo C-terminal marcadas con 6xHis. La DHFR (dihidrofolato reductasa) mejora la antigenicidad y la estabilidad y se recomienda para las proteínas poco expresadas o péptidos cortos propensos a la proteólisis. Como la propia DHFR muestra poco inmunogenicidad en ratones y ratas, las proteínas de fusión de DHFR son ideales para el cribado de epítopos.
Principle
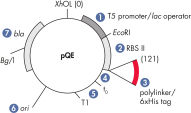
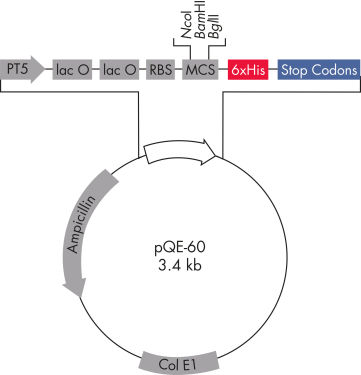
Los vectores pQE de QIAexpress pQE combinan un potente promotor del fago T5 (reconocido por la ARN polimerasa de E. coli) con un módulo de represión de doble operador lac para proporcionar una expresión estrictamente regulada y de alto nivel de las proteínas recombinantes en E. coli (consulta la figura « Vector pQE de QIAexpress»). La síntesis de proteínas se bloquea de forma efectiva en presencia de altos niveles del represor lac y mejora la estabilidad de preparados citotóxicos. Los vectores pQE permiten la colocación del marcado 6xHis ya sea en el extremo N-terminal o en el C-terminal de la proteína recombinante.
| Elemento | Descripción |
| Elemento optimizado del promotor/operador | Consta del promotor del fago T5 y dos secuencias del operador lac, que aumentan la probabilidad de la unión del represor lac y garantizan una represión eficiente del potente promotor T5 |
| Sitio de unión ribosómica sintética RBSII | Para una traslación eficiente |
| Secuencia de codificación marcada con 6xHis | 5' o 3’ a la región de clonación del policonector |
| Codones de parada traslacionales | En todos los marcos de lectura, para la preparación práctica de preparados de expresión |
| Dos potentes terminadores transcripcionales | t0 del fago lambda y T1 del operón rrnB de E. coli, para evitar la transcripción de lectura y garantizar la estabilidad de la construcción de expresión |
| Origen de replicación de ColE1 | A partir de pBR322 |
| Gen de la betalactamasa (bla) | Confiere resistencia a la ampicilina |
See figures
Procedure
Los insertos que codifican proteínas de interés se clonan en construcciones adecuadas (para obtener información detallada, consulte El Manual QIAexpressionist) y se transformaron en una cepa de E.coli para la expresión. La expresión se induce mediante la adición de IPTG. Los preparados del vector pQE TriSystem se pueden transformar en E.coli, utilizarse como vector lanzadera para la expresión de proteína recombinante en células de insecto o transmitirse a células de mamíferos.
Applications
El sistema Expression QIAexpress proporciona una expresión de proteínas de alto nivel adecuado para
muchas aplicaciones, entre las que se incluyen:
- Purificación de proteínas funcionales y conformacionalmente activas
- Purificación en condiciones desnaturalizantes para la producción de anticuerpos
- Cristalización para la determinación de la estructura tridimensional
- Ensayos relacionados con interacciones proteína-proteína y proteína-ADN
Supporting data and figures
Vector de QIAeexpress pQE.
Specifications
| Features | Specifications |
|---|---|
| Expression species | E. coli |
| In-frame cloning necessary | Sí |
| N- or C-terminal tag | Marcador de extremo C-terminal |
| Expression | In vivo |
| Special features | Basado en el sistema de transcripción-traducción del promotor T5 |
| Tag | Etiqueta 6xHis |
| Tag removal sequence | No |
| All three reading frames provided | Sí |