Type-it Fast SNP Probe PCR Kit
应用TaqMan或TaqMan MGB探针进行准确可靠的SNP基因分型
应用TaqMan或TaqMan MGB探针进行准确可靠的SNP基因分型
✓ 全天候自动处理在线订单
✓ 博学专业的产品和技术支持
✓ 快速可靠的(再)订购
Cat. No. / ID: 206045
✓ 全天候自动处理在线订单
✓ 博学专业的产品和技术支持
✓ 快速可靠的(再)订购
Type-it Fast SNP Probe PCR Kit确保持续获得高度准确的SNP基因分型。
Type-it Fast SNP Genotyping PCR Master Mix基于高度特异性的HotStarTaq Plus DNA Polymerase和新研发的SNP基因分型PCR缓冲液体系,可进行高度特异性的探针结合和持续强的荧光信号。与其他商业化SNP基因分型预混液相比,可获得更广泛、更清晰的等位基因群分离(参见" Highly specific probe binding")。
对于有挑战性的靶点或难处理的SNP位点,Type-it Fast SNP Probe PCR Kit提供可靠的SNP基因分型,可完全分开等位基因并使相同的等位基因紧密成群,产生较高的信号率(参见" Successful automated allele calling")。Type-it Fast SNP Probe PCR Kit优于其他供应商的试剂盒,保证等位基因紧密成群,即使1 ng的DNA模板(参见 " Reliable SNP genotyping with small amounts of template")。
Type-it Fast SNP Probe PCR Kit采用方便的预混液规格,含有高度特异性的HotStarTaq Plus Polymerase和独特的SNP基因分型缓冲液体系,可快速、高效的扩增。独特的试剂盒组分可使等位基因紧密成群,完全分开确保高信号率,获得可重复的、准确的结果(见表)。
预混液中提供的HotStarTaq Plus DNA Polymerase确保高特异性扩增,即使处理低模板量。HotStarTaq Plus DNA Polymerase处于非活化状态,在环境温度下没有聚合酶活性。这避免了在低温下引物错搭产物和引物二聚体的形成。
预混液中含有的Type-it Fast SNP PCR Buffer专用于通过序列特异性5'核酸酶探针进行快速循环的SNP基因分型。Type-it Fast SNP PCR Buffer的独特组分确保非常严格和特异性的等位基因特异性探针的结合。改变探针的熔解行为可形成更窄的探针熔解温度窗口。基于原始的QIAGEN PCR Buffer,这种创新的缓冲液使每个PCR循环的退火步骤都有较高比率的特异性引物结合。KCl和(NH4)2SO4独特比例结合,使PCR缓冲液与常规PCR缓冲液相比,可在更宽范围的退火温度和Mg2+浓度下提供严格的引物退火条件。极大减少通过改变退火温度或Mg2+浓度的PCR优化过程,有时甚至不需要。
Type-it Fast SNP PCR缓冲液中的添加剂,如Q-Solution,可提供反应条件,用于难处理基因组区域和SNP位点的扩增。创新的Q-Bond技术(用于快速循环),可更快获得SNP基因分型结果(参见" Mechanism of fast cycling during annealing")。预混液中含有的ROX染料,其浓度使其能在Applied Biosystems的多种仪器上进行SNP基因分型。而且,ROX染料也能在不需要ROX作为阴性参照染料的仪器上使用(见表)。
| 试剂盒组分 | 特征 |
|---|---|
| 2x Master Mix format* | 用于配合TaqMan MGB探针进行SNP基因分型。能与多种循环仪配合使用,用于SNP基因分型 |
| HotStarTaq Plus DNA Polymerase | 室温下进行快速、简单的反应体系构建。即使使用少量模板,也能高度特异性扩增 |
| Type-it Fast SNP Probe PCR Buffer | 广泛分离基因群。紧密的等位基因群,高等位基因信号率。探针结合具有更高特异性。使用Q-Bond Molecule,快速循环 |
| Q-Solution | 对于高GC含量的SNP位点,在散点图分析中具有更紧密的集群和更强信号。可进一步提高不理想的等位基因信号 |
Type-it Fast SNP Probe PCR Kit使用商业化的SNP基因分型检测进行了功能验证,能与TaqMan MGB Probes配合使用,也可配合用户自行设计的探针法检测,可使用TaqMan MGB、TaqMan或其他双标记探针。此试剂盒包括简化的、预优化的操作流程,用于快速、可靠的分析。
Type-it Fast SNP Probe PCR Master Mix提供的反应条件,配合序列特异性探针进行快速PCR循环。缓冲液包含专有化的Q-Bond Molecule,可在标准循环仪或具有快速升温速率的快速循环仪上缩短循环时间。快速循环步骤可减少至多40%的PCR运行时间,提高通量(参见" Genotyping workflow")。
该试剂盒采用即用型、预优化预混液形式,具有更大便利性。使用预混液可节约时间,简化反应体系构建的手工操作,通过消除移液误差和污染的可能来源,提高可重复性,尽量减少移液步骤,无需繁琐的计算。预混液中含有的HotStarTaq Plus DNA Polymerase 95°C,5分钟被激活,可在热循环程序中简单设置。使用预混液进行室温下的反应体系构建快速、简单。
Type-it Fast SNP Probe PCR Kit经优化,可在多种合适的标准和快速升温的循环仪上进行SNP PCR分析,光学PCR孔板配合real-time PCR仪可用于荧光板阅读分析(见表)。
| 循环仪 | 模式 |
|---|---|
| Real-time PCR cyclers | QIAGEN: Rotor-Gene Q ABI PRISM 7900 (all series) Applied Biosystems 7500 (all series) Applied Biosystems 7300 ABI PRISM 7700 ABI PRISM 7000 ABI StepOne and StepOnePlus Bio-Rad: iCycler iQ Roche: LightCycler 480 Stratagene: Mx3000P and Mx3005P |
| Standard cyclers | 采用标准升温模式的所有循环仪(如GeneAmp 9700) 采用快速升温模式的所有循环仪(如GeneAmp 9800) |
Type-it Fast SNP Probe PCR Kit应用TaqMan MGB探针检测SNPs,可用于多个研究领域,包括:
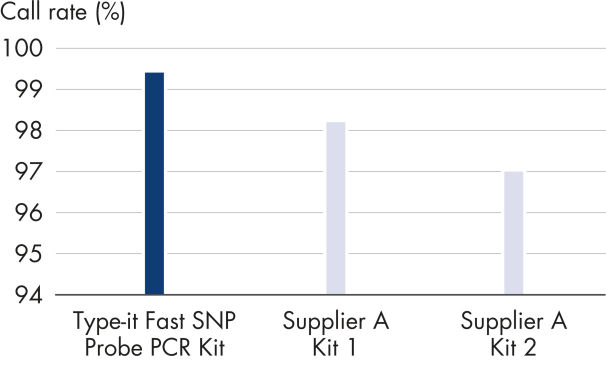
Average automated allele call rates for a panel of different DNAs were analyzed using 6 different TaqMan® MGB-based SNP genotyping assays and 1 ng of genomic DNA from each of the 60 samples. PCR was performed with the indicated products in 384-well plates in a 5 μl reaction volume. Allelic discrimination plate read was performed on an Applied Biosystems 7900HT instrument. The Type-it Fast SNP Probe PCR Kit consistently resulted in the highest call rates and the lowest error rates.
| Features | Specifications |
|---|---|
| Applications | Probe-based SNP Genotyping |
| Product use | Functionally validated and developed for reliable SNP Genotyping |
| Mastermix | Yes |
| Real-time or endpoint | Both |
| With or without ROX | ROXincludedinMasterMix |
| With/without hotstart | With |
| Sample/target type | GenomicDNA |
| Enzyme activity | 5'->3'Exonucleaseactivity |
| Sequence specific Probe | TaqMan®GenotypingAssays,TaqMan®orTaqManMGB®probes |
| Reaction type | PCRamplification |