✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Cat. No. / ID: 180492
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Ultra-efficient chemistries maximize performance of sub-nanogram samples
- Flexible protocol provides a comprehensive solution for 10 pg−100 ng DNA input
- Compatible with a wide range of sample types, including cfDNA, FFPE, ancient DNA and ChIP-seq
- Unique Dual Index Adapters available for multiplexing up to 384 samples
Product Details
The QIAseq Ultralow Input Library Kit incorporates new, ultra-efficient end-repair and ligation chemistries into a complete and streamlined solution. These kits allow the generation of high-quality libraries from limited and damaged DNA sample types using an efficient, 2.5-hour workflow – starting from 10 pg–100 ng fragmented DNA for a robust library preparation suitable for a wide range of research applications.
Want to try this solution for the first time? Request a quote for a trial kit.
Performance
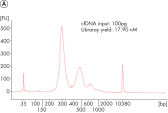
Maximum yield from minimal input
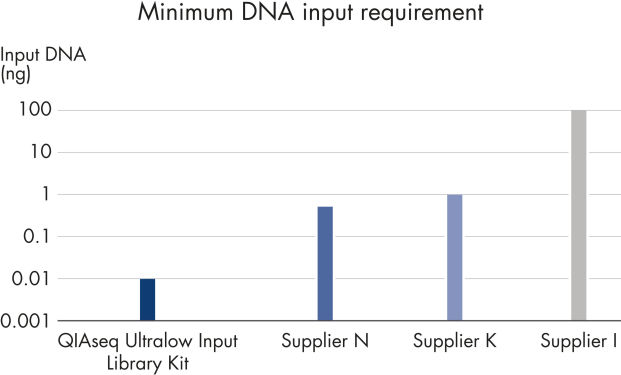
Confidently process samples from 10 pg to 100 ng DNA input. The new, highly efficient chemistries in QIAseq Ultralow Input Kits eliminate the issue of performance drop-off below 1 ng of input DNA. Novel enzymes and buffer formulations maximize the performance of sub-nanogram samples while also enabling the flexibility to process samples with up to 100 ng input DNA.
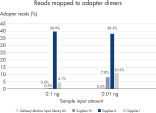
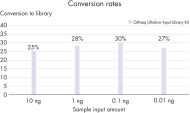
Highly efficient and consistent conversion rates
QIAseq Ultralow Input Library Kits enable superior library conversion rates (>30%) from samples below 1 ng of input using newly optimized chemistry formulations. These consistent conversion rates of genomic information to adapter-ligated library molecules cover a 1000-fold range – from 10 pg to 10 ng of DNA input – and set an industry-leading benchmark. The low percentage of adapter dimers minimizes library contamination compared to the competition.
Robust solution for challenging samples
QIAseq Ultralow Input chemistry is ideal for: low-input whole genome sequencing; cfDNA for liquid biopsy applications; and limited sources of DNA from FFPE tissue, laser capture microdissection (LCM) and fine needle aspirations (FNA).
Principle
Next-generation sequencing (NGS) is a driving force for many new and exciting applications, including cancer research, stem cell research, metagenomics, population genetics and medical research. While NGS technology is continuously improving, library preparation remains a process bottleneck for many labs and a limiting factor in the types of samples that can successfully generate NGS data.
As a result, capturing the maximum possible genomic information in an NGS library is both critical and challenging, when starting from limited or damaged DNA samples. Standard library preparation methods that apply to many sample types – such as circulating cell-free DNA, FFPE DNA, ancient DNA and ChIP-seq DNA – can result in quality-related problems. These problems limit the utility of the data generated, from low library yield or conversion rates to high percentages of reads derived from adapter dimer molecules.
Procedure
Library construction
QIAseq Ultralow Input Library Kits start with double-stranded DNA that has been fragmented enzymatically, chemically, mechanically or naturally. Kits use an optimized end-polishing reaction and a new Ultralow Input Ligation formulation, along with QIAGEN’s proprietary HiFi PCR Master Mix. This combination maximizes the conversion rate of sample DNA into the NGS library, while efficiently and evenly amplifying even high and low GC content regions of the genome. This protocol enables the highest possible yield sequencing library, free of adapter dimer contamination – starting from a little as 10−100 pg* of DNA input. Due to the kit’s flexible protocol, the same kit can also be used for higher DNA input amounts, including PCR-free library preparations from as little as 10 pg* of DNA input.
*Note that the genomic complexity necessary for a given experiment will vary depending on the genome size of the organism and the fraction of that genome included in the target region of interest.>
Reaction cleanup
Following adapter ligation and library amplification steps, reaction cleanup and removal of residual adapter dimers can be achieved by using Agencourt AMPure XP beads – which enable easy automation on various high throughput automation platforms. >
Optional amplification of library DNA
PCR-based library amplification is normally required if the input DNA amount is below 100 ng. This protocol is optional, and enables high-fidelity amplification of the DNA library using the QIAseq HiFi PCR Master Mix that is included in the kit.
Contents and accessories
Dual-barcoded, plate-format adapters are included with the 96-reaction size QIAseq Ultralow Input Library Kit. Each well in the 96-plex adapter plate contains a single-use adapter consisting of a unique combination of two eight-nucleotide identification barcodes. By combining one i5 barcode and one i7 barcode in each ready-to-use adapter, the 96 reaction QIAseq Ultralow Input Kits support up to 96-plex pooling of libraries prior to sequencing.
While adapters are not included in the 12-reaction size kit, the QIAseq UDI Y-Adapter Kit (24), cat. no. 180310, containing dual bar-coded Unique Dual Sample Index Y-adapters (plate-format for 24 samples) can be ordered separately.
Applications
QIAGEN QIAseq Ultralow Input Library Kits have been designed to be the definitive solution for generating high quality libraries from even very challenging NGS samples. Intended for NGS researchers who seek a single library prep kit compatible with a wide range of ultra-low, low and standard input fragmented DNA – QIAseq Ultralow Input Library Kits enable new insights by maximizing performance, particularly from limited and damaged DNA sample types. The streamlined, 2.5-hour protocol for generating libraries from fragmented DNA using QIAseq Ultralow Input Library Kits also enables straightforward automation on different liquid-handling platforms.
Supporting data and figures
Minimum DNA input requirements for a selection of currently available library preparation products.