✓ オンライン注文による24時間年中無休の自動処理システム
✓ 知識豊富で専門的な製品&テクニカルサポート
✓ 迅速で信頼性の高い(再)注文
REPLI-g Mitochondrial DNA Kit (25)
カタログ番号 / ID. 151023
✓ オンライン注文による24時間年中無休の自動処理システム
✓ 知識豊富で専門的な製品&テクニカルサポート
✓ 迅速で信頼性の高い(再)注文
特徴
- 様々なスタートサンプルから高感度な増幅
- ヒトおよびヒト以外のミトコンドリア全ゲノムの均一な増幅
- 時間のかかるミトコンドリアDNAの精製が不要
- 損傷核DNAを持つサンプルからも有用な結果
- Blood cardや毛髪からのDNA増幅も可能
製品詳細
REPLI-g Mitochondrial DNA Kitを用いることにより、ミトコンドリアDNAを事前に精製することなく、トータルDNAから選択的にミトコンドリアDNAを増幅できます。本キットにはDNAポリメラーゼ、バッファー、試薬が含まれ、トータルDNA中の微量のミトコンドリアDNAサンプルから特異的かつ均一なミトコンドリアの全ゲノムDNA増幅を実現します。ヒト以外の生物種のミトコンドリアDNA を選択的に増幅されたい場合は、キットに含まれているREPLI-g Human mt Primer Mixではなく、適切なPrimer Mix をご用意ください。REPLI-g Mitochondrial DNA Kit は、Multiple Displacement Amplification(MDA)テクノロジーにより、簡単な操作で核DNAのコンタミがほとんどないミトコンドリアDNAの全ゲノム増幅を実現するため、時間のかかるミトコンドリアDNA精製の必要がなく、ダウンストリームアッセイの感度を向上できます。
パフォーマンス
原理
ミトコンドリアDNAを使用する際の1つの制限は、核DNAからミトコンドリアDNAを精製しなければならないことでした(特にミトコンドリアDNAマーカーのアッセイ感度を高める必要がある場合に重要)。この精製操作には時間のかかる多くのステップが必要であり、ミトコンドリアDNAをかなり損失することになります。REPLI-g Mitochondrial DNA KitはトータルDNAサンプル中のミトコンドリアDNA配列を増幅することで、この問題を解決しています。
REPLI-g Mitochondrial DNAテクノロジーは、ミトコンドリアのゲノムDNA領域の迅速で均一な全ゲノム増幅を実現します。この方法は、伸長性が高いユニークなDNAポリメラーゼを用いて等温でゲノム増幅ができるMultiple Displacement Amplification(MDA)テクノロジーを利用しています。REPLI-g DNA Polymeraseは高い伸長性と相補鎖置換能により、DNAテンプレートから解離せずに最高100 kbまでの増幅ができます。 そのため、PCRベースの全ゲノム増幅法と比べミスマッチ配列と遺伝子座を最小限に抑えます。
操作手順
アプリケーション
REPLI-gテクノロジーで増幅したミトコンドリアDNAは次のような様々なダウンストリームアプリケーションに使用できます。
- ジェノタイピング(SNP、欠失、挿入等)
- エンドポイントPCRおよび定量リアルタイムPCR
- シークエンシング解析
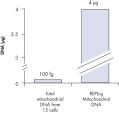
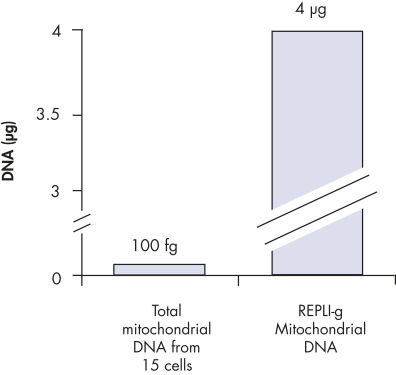
裏付けデータと数値
Enrichment of mitochondrial DNA.
仕様
| 特徴 | 仕様 |
|---|---|
| Amplification | Whole genomic DNA |
| Samples per run (throughput) | Mid |
| Denaturation step | Heat |
| Maximum input volume | 10 µl template DNA |
| Minimal pipetting volume needed | 1 µl |
| Reaction volume | 50 µl |
| Reaction time | ~8 hours (overnight) |
| Quality assessment | No |
| Applications | Genotyping, sequencing, RFLP |
| Starting amount of DNA | ~10 ng purified total DNA |
| Starting material | Genomic human DNA |
| Technology | Multiple Displacement Amplification (MDA) |
| Yield | ~4 µg |