- Experiment Configurator
- Discovery & Translational Research
- Diagnostics & Clinical Research
- Human ID & Forensics
- Next-Generation Sequencing
- Instruments & Automation
- Informatics & Data
- Services
- OEM by QIAGEN
- Top Sellers
- New Solutions
Enzymes for molecular biology
Explore high-quality enzymes; now available as individual products.
Products and tools for your targets
Explore targets and pathways in their scientific context, find and customize products to study them, analyze data and plan follow-up studies – all in GeneGlobe.
QuantiFERON-TB Gold Plus
Detect TB infection with confidence.

✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Cat. No. / ID: 250031
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Four nanoplates for different application needs
- Up to 8500 or 26,000 partitions per well
- SBS format
Product Details
The QIAcuity Nanoplates are microfluidic digital PCR plates that enable running 8, 24 or 96 samples with up to 8500 or 26,000 partitions per well. All four nanoplates are designed to run on the QIAcuity digital PCR instruments.
The nanoplates can only be used with the QIAcuity Digital PCR System. Use the dedicated QIAcuity Nanoplate Adapter when performing automated liquid handling and PCR setup in a QIAcuity Nanoplate using the QIAgility. Once done, load the plate onto the QIAcuity Digital PCR System for the dPCR reaction.
Would you like to learn more about the product and be contacted by one of our dPCR specialists? Sign in here, and we will get in touch with you shortly.
Performance
These are specially designed plates used for the digital PCR reactions in the QIAcuity instruments. QIAGEN offers four nanoplate types, all SBS format, but with different specifications for different application needs.
| Type | Frame color | Specifications | Applications |
| Nanoplate 26K 8-well | Light blue |
8-well x approx. 26,000 partitions 40 µl dPCR reaction per well |
Rare mutation detection, liquid biopsy, pathogen detection, etc. |
| Nanoplate 26K 24-well | Blue |
24-well x approx. 26,000 partitions 40 µl dPCR reaction per well |
|
| Nanoplate 8.5K 24-well | White |
24-well x approx. 8500 partitions 12 µl dPCR reaction per well |
Copy number variation analysis, gene expression analysis, NGS library quantification, genome edit detection, etc. |
| Nanoplate 8.5K 96-well | Gray |
96-well x approx. 8500 partitions 12 µl dPCR reaction per well |
Principle
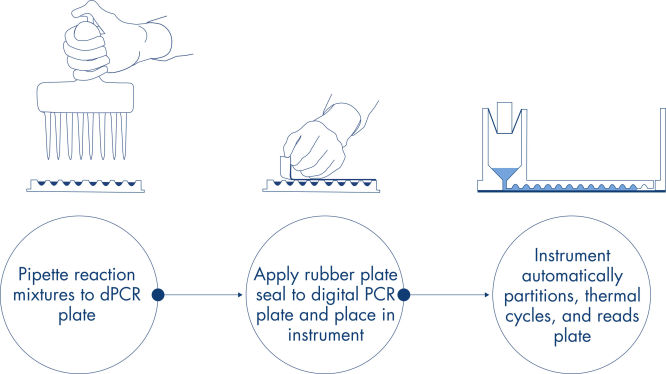
In just 3 simple steps, you can have the dPCR result you want in under 2 hours: pipette and load, run the experiment, analyze results. The principle of the dPCR reaction in the nanoplates is described here.
Procedure
Just like in qPCR experiments, sample preparation includes the transfer of master mix, probes and primers to a 8-, 24- or 96-well nanoplate, followed by the addition of samples. The system integrates partitioning, thermocycling and imaging into a single fully automated instrument that takes users from the sample to result in under 2 hours. One can perform analysis on the Suite Software, providing the concentration in copies per microliter of your target sequence as well as for quality control such as positive samples or NTC. This analysis can also be extended to remote computers within the same local area network (LAN).
Applications
The QIAcuity Nanoplates, in combination with the QIAcuity Digital PCR System and the QIAcuity PCR kits, enable digital PCR applications, including:
- Rare mutation detection
- Copy number variation analysis
- Gene expression analysis
- Pathogen detection
- Genotyping
- miRNA research
- Cell and gene therapy
- Residual DNA quantification
- Waste water monitoring
Supporting data and figures
A simple and rapid plate-based workflow
Resources
Version 2.1
Version 2.1
Version 2.1.8
Version 2.1.8
Version 2.5.0.0
Version 2.5.0.1
Version 2.5.0.1
The QIAcuity Software Suite enables the user to set up plates, analyze results, and monitor the status of runs in real time. This software is also used for the configuration of the system and provides access to the QIAcuity user management. The QIAcuity Software Suite is designed to be installed on a Windows PC that is connected to one or more QIAcuity instruments. For this configuration, the QIAcuity instrument needs to be connected to a network through Ethernet. Alternatively, a direct cable connection between the QIAcuity and the notebook where the QIAcuity Software Suite is running needs to be established. When connected to a network, up to 10 users may access the QIAcuity Software Suite via a browser installed on the client PC (Windows or Mac).
The new version 2.5 of the QIAcuity Software Suite supports assay development by providing an essential temperature gradient functionality. It also offers a new feature that provides calculation of initial concentration of samples by using various dilution factors. In addition, concentration units may be converted into various pre-defined or user-defined units. The new software also offers an integrity value and concentration value per group for up to 5-plex added to the multiple occupancy CSV file export, for example, for the evaluation of AAV (adeno-associated virus) assays and for drop-off assays.
In this new version the initial loading time and the time for recalculation of 1D/2D scatterplot and for signal map image has been reduced, leading to a much faster performance.
Detailed information about the QIAcuity Software Suite version 2.5 is available in the Release Note, which can also be downloaded under section “Software Release Notes”.
Note: The latest Software Suite version 2.5 is only compatible with the latest Control Software (CSW) version 2.5. If only one software component is updated, no connection between the Software Suite and the CSW can be established.
Important: Please follow the instructions provided in the QIAcuity User Manual for software version 2.5 and in the Release Note.
It is strongly recommended to update the QIAcuity Software Suite first before proceeding with the QIAcuity Control Software!
Please contact QIAGEN Technical Services if you are unsure and require technical support.
SHA1 checksum: D1690226299A75E077FA37C420A970FC71D56CBF
The following QIAcuity Software Suite Volume Precision Factor (VPF) patches have been released to enable compatibility with VPF file version 6.0 or higher for QIAcuity Software Suite version 1.2.18. As the abovementioned QIAcuity Software Suite versions do not allow loading a VPF zip file including more than 20 different individual VPF files, a patch is needed to allow the software for loading VPF zip files version 6.0 or higher. This technical limitation is solved from QIAcuity Software Suite version 2.1.8.23 onwards.
If you are using the QIAcuity Software Suite version 1.2.18, please consider updating your Software Suite and the Instrument Control Software (CSW) to the most recent version. After updating to the latest software version, no patching is needed.
If you are not able to update your QIAcuity Software Suite, please install the QIAcuity Software Suite VPF patch for QIAcuity Software Suite version 1.2.18 before loading VPF file version 6 or higher. Additional information and instructions may be found on the Release Note: QIAcuity Software Suite Volume Precision Factor (VPF) Patches.
The following QIAcuity Software Suite Volume Precision Factor (VPF) patches have been released to enable compatibility with VPF file version 6.0 or higher for QIAcuity Software Suite versions 2.0.20. As the abovementioned QIAcuity Software Suite versions do not allow loading a VPF zip file including more than 20 different individual VPF files, a patch is needed to allow the software for loading VPF zip files version 6.0 or higher. This technical limitation is solved from QIAcuity Software Suite version 2.1.8.23 onwards.
If you are using the QIAcuity Software Suite version 2.0.20, please consider updating your Software Suite and the Instrument Control Software (CSW) to the most recent version. After updating to the latest software version, no patching is needed.
If you are not able to update your QIAcuity Software Suite, please install the QIAcuity Software Suite VPF patch for QIAcuity Software Suite version 2.0.20 before loading VPF file version 6 or higher. Additional information and instructions may be found on the Release Note: QIAcuity Software Suite Volume Precision Factor (VPF) Patches.
The following QIAcuity Software Suite Volume Precision Factor (VPF) patches have been released to enable compatibility with VPF file version 6.0 or higher for QIAcuity Software Suite version 2.1.7.182. As the abovementioned QIAcuity Software Suite versions do not allow loading a VPF zip file including more than 20 different individual VPF files, a patch is needed to allow the software for loading VPF zip files version 6.0 or higher. This technical limitation is solved from QIAcuity Software Suite version 2.1.8.23 onwards.
If you are using the QIAcuity Software Suite version 2.1.7.182, please consider updating your Software Suite and the Instrument Control Software (CSW) to the most recent version. After updating to the latest software version, no patching is needed.
If you are not able to update your QIAcuity Software Suite, please install the QIAcuity Software Suite VPF patch for QIAcuity Software Suite version 2.1.7.182 before loading VPF file version 6 or higher. Additional information and instructions may be found on the Release Note: QIAcuity Software Suite Volume Precision Factor (VPF) Patches.
The following QIAcuity Software Suite Volume Precision Factor (VPF) patches have been released to enable compatibility with VPF file version 6.0 or higher for QIAcuity Software Suite version 2.1.8.20. As the abovementioned QIAcuity Software Suite versions do not allow loading a VPF zip file including more than 20 different individual VPF files, a patch is needed to allow the software for loading VPF zip files version 6.0 or higher. This technical limitation is solved from QIAcuity Software Suite version 2.1.8.23 onwards.
If you are using the QIAcuity Software Suite version 2.1.8.20, please consider updating your Software Suite and the Instrument Control Software (CSW) to the most recent version. After updating to the latest software version, no patching is needed.
If you are not able to update your QIAcuity Software Suite, please install the QIAcuity Software Suite VPF patch for QIAcuity Software Suite version 2.1.8.20 before loading VPF file version 6 or higher. Additional information and instructions may be found on the Release Note: QIAcuity Software Suite Volume Precision Factor (VPF) Patches.
The QIAcuity backup and restore scripts are a standalone solution to backup all relevant user data of the QIAGEN Software Suite for disaster recovery and restore the data on a new or existing QIAGEN Software Suite installation. The following QIAcuity Software Suite versions are supported: 2.0, 2.1.7, 2.1.8, and 2.2. The backup can be conducted manually or automated by using of the Windows task scheduler.
Note: Windows Admin permission is needed to setup and perform an automated backup and for manual backup and restore.
Please read the QIAcuity Software Suite Backup and Restore Scripts document for more information and instructions how to use the scripts.
The QIAcuity backup and restore scripts are a standalone solution to backup all relevant user data of the QIAGEN Software Suite for disaster recovery and to restore the data on a new or existing QIAGEN Software Suite installation. The backup supports the QIAcuity Software Suite version 2.5 or higher and can be conducted manually or automatically by using of the Windows Task Scheduler.
Note: Windows Admin permission is needed to setup and perform an automated backup and for manual backup and restore.
For backup and restore of Software versions lower than 2.5, please refer to the QIAcuity Software Suite Backup and Restore Scripts for version 2.0, 2.1.7, 2.1.8 and 2.2.
Note: Windows Admin permission is needed to setup and perform an automated backup and for manual backup and restore.
Please read the QIAcuity Software Suite Backup and Restore Scripts document for more information and instructions how to use the scripts.
Version 2.5
The QIAcuity Control Software is an integral part of the QIAcuity instrument. It offers a GUI (graphical user interface) for basic functionalities such as plate setup, changing the order of plates to be processed, and monitoring the status of runs in real time. After a run is completed, the data are stored on the instrument’s memory and are sent to the connected QIAcuity Software Suite for analysis.
The new version 2.5 of the QIAcuity CSW offers a configurable auto logoff times which enables users to turn off or define logoff times per instrument. In addition the new version supports assay development by providing an essential temperature gradient functionality.
Furthermore improvements were implemented, for example, for the accuracy of time estimation for various software steps.
Detailed information about the QIAcuity Control Software version 2.5 is available in the Release Note, which can also be downloaded under section Software Release Notes.
Note: The latest CSW version 2.5 is only compatible with the Software Suite version 2.5. If only one software component is updated, no connection between the Software Suite and the CSW can be established.
Important: Please follow the instructions provided in the QIAcuity User Manual for software version 2.5 and in the Release Note.
It is strongly recommended to update the QIAcuity Software Suite first before proceeding with the QIAcuity Control Software!
Note: After clicking reboot during CSW upgrade or change of Suite connection, the login screen may appear for short period. Please ignore it and wait for the QIAcuity instrument to shut down and restart itself.
Please contact QIAGEN Technical Services if you are unsure and require technical support.
Version 8.0
The Volume Precision Factor (VPF) offers a unique feature to secure precision of concentration results obtained from a QIAcuity dPCR run.
In general, Nanoplates provide partitions of fixed sizes that enable a very precise way of sample concentration calculation. Potential variation of partition sizes in Nanoplate batches, caused by different microstructure molding forms, can be addressed by applying the batch specific VPF. Furthermore, the VPF includes well-specific volume information and therefore further increases precision of concentration calculation in each well of the Nanoplates.
Important note:
The Volume Precision Factor file version 8.0 is compatible with QIAcuity Software Suite version 2.1.8.23 or higher. All lower versions, namely 1.2.18, 2.0.20, 2.1.7.182, and 2.1.8.20, require a QIAcuity Software Suite patch to be installed prior the upload of VPF version 8.0. Please read the Release Note for QIAcuity Software Suite Volume Precision Factor (VPF) Patches for more information.
If you are using QIAcuity Software Suite versions older than 2.1.8.23, please consider updating your Software Suite and the Instrument Control Software (CSW) to the most recent version prior the upload of VPF version 8.0. After updating to the latest QIAcuity Software Suite software version, no patching is needed. If you are not able to update your QIAcuity Software Suite, please run the patch for your Software Suite version following the instructions provided in the Release Note for QIAcuity Software Suite Volume Precision Factor (VPF) Patches.
After downloading and updating the VPF file within the QIAcuity Software Suite, the VPF is applied automatically to the analysis of a corresponding Nanoplate batch. The VPF file includes information from all available microstructure molding forms and connected Nanoplate batches. It will be stored on the PC where the QIAcuity Software Suite is installed.
Version 2.1.8
The QIAcuity Software Suite enables the user to set up plates, analyze results, and monitor the status of runs in real time. This software is also used for the configuration of the system and provides access to the QIAcuity user management. The QIAcuity Software Suite is designed to be installed on a Windows PC that is connected to one or more QIAcuity instruments. For this configuration, the QIAcuity instrument needs to be connected to a network through Ethernet. Alternatively, a direct cable connection between the QIAcuity and the notebook where the QIAcuity Software Suite is running needs to be established. When connected to a network, up to 10 users may access the QIAcuity Software Suite via a browser installed on the client PC (Windows or Mac).
The new version 2.1.8 of the QIAcuity software (QIAcuity Software Suite version 2.1.8 and QIAcuity CSW version 2.1.8.) offers bug fixes for repeated network connection issues and improved error handling. In addition, it fixes issues seen with latest versions of Microsoft Edge and Google Chrome browsers as well as issues seen with read-only plates after a software update.
Detailed information about the QIAcuity Software Suite version 2.1.8 is available in the Release Note, which can also be downloaded under section “Software Release Notes”.
Important: Please ensure that updating to QIAcuity Software Suite version 2.1.8 is performed by the same Windows admin user using the same Windows login name that installed the previous QIAcuity Software Suite version. In case you cannot use the same Windows login name or the software update resulted in the browser error message "Can't reach this page" please contact our Technical Services for additional instructions.
Note: The latest Software Suite version 2.1.8 is only compatible with the latest Control Software (CSW) version 2.1.8. If only one software component is updated, no connection between the Software Suite and the CSW can be established.
Important: Please follow the instructions provided in the QIAcuity User Manual for software version 2.1 and in the Release Note. It is strongly recommended to update the QIAcuity Software Suite first before proceeding with the QIAcuity Control Software!
Please contact QIAGEN Technical Services if you are unsure and require technical support.
SHA1 checksum: DE2416D926BB98D82A332E7B25EF66728F120DDA
Version 2.2
The QIAcuity Software Suite enables the user to set up plates, analyze results, and monitor the status of runs in real time. This software is also used for the configuration of the system and provides access to the QIAcuity user management. The QIAcuity Software Suite is designed to be installed on a Windows PC that is connected to one or more QIAcuity instruments. For this configuration, the QIAcuity instrument needs to be connected to a network through Ethernet. Alternatively, a direct cable connection between the QIAcuity and the notebook where the QIAcuity Software Suite is running needs to be established. When connected to a network, up to 10 users may access the QIAcuity Software Suite via a browser installed on the client PC (Windows or Mac).
The new version 2.2 of the QIAcuity software (QIAcuity Software Suite v 2.2 and QIAcuity CSW v 2.2.) offers improvements for users working under GMP by adding a user ID validation during the report signing and the addition of timezone offset stamp for audit trail entries and for result report data. Furthermore, the addition of a standard deviation and coefficient of variance calculation in percentage of mean concentration calculation for replicates where implemented. In addition, the instrument camera stability was improved and an internal validation method was implemented.
Detailed information about the QIAcuity Software Suite version 2.2 is available in the Release Note, which can also be downloaded under section “Software Release Notes”.
Note: The latest Software Suite version 2.2 is only compatible with the latest Control Software (CSW) version 2.2. If only one software component is updated, no connection between the Software Suite and the CSW can be established.
Important: Please follow the instructions provided in the QIAcuity User Manual for software version 2.2 and in the Release Note.
It is strongly recommended to update the QIAcuity Software Suite first before proceeding with the QIAcuity Control Software.
Please contact QIAGEN Technical Services if you are unsure and require technical support.
SHA1 checksum: 63A1689F2EB557809F418D159DA438FDC8B327A3
Version 2.1.8
The QIAcuity Control Software (CSW) is an integral part of the QIAcuity instrument. It offers a GUI (graphical user interface) for basic functionalities such as plate setup, changing the order of plates to be processed, and monitoring the status of runs in real time. After a run is completed, the data are stored on the instrument’s memory and are sent to the connected QIAcuity Software Suite for analysis.
The new version 2.1.8 of the QIAcuity software (QIAcuity Software Suite version 2.1.8 and QIAcuity CSW version 2.1.8) offers bug fixes for repeated network connection issues and improved error handling. In addition, it fixes issues seen with latest versions of Microsoft Edge and Google Chrome browsers as well as issues seen with read-only plates after a software update.
Detailed information about the QIAcuity Control Software version 2.1.8 is available in the Release Note, which can also be downloaded under section “Software Release Notes”.
Note: The latest CSW version 2.1.8 is only compatible with the latest Software Suite version 2.1.8. If only one software component is updated, no connection between the Software Suite and the CSW can be established.
Important: Please follow the instructions provided in the QIAcuity User Manual for software version 2.1 and in the Release Note. It is strongly recommended to update the QIAcuity Software Suite first before proceeding with the QIAcuity Control Software!
Please contact QIAGEN Technical Services if you are unsure and require technical support.
SHA1 checksum: D3982B419D7C4CF39FBDDDAA4C0D351B39398278
Version 1.2
A newer version of the Software Suite is available. Please use this version for the update of older plates if required.
The QIAcuity Software Suite 1.2 is designed to be installed on a Windows PC that is connected to one or more QIAcuity instruments. The QIAcuity Software Suite enables the user to set up plates, analyze results, and monitor the status of runs in real time. For this configuration, the QIAcuity instrument needs to be connected to a network through Ethernet. Alternatively, a direct cable connection between the QIAcuity and the notebook where the QIAcuity Software Suite is running needs to be established. When connected to a network, up to 10 users may access the QIAcuity Software Suite via a browser installed on the client PC (Windows or Mac).
The following browsers are supported in the QIAcuity Software Suite:
-Mozilla Firefox (version 64.0.2 or higher)
-Microsoft Edge (version 44.17763.1.0 or higher)
-Google Chrome (version 71.0.3578.98 or higher)
The new QIAcuity Software Suite 1.2 offers a functionality that enables users of the QIAcuity Software 1.1.3 to upgrade to the new version while keeping the library of previously stored plate runs.
Note: If you have exported plates from QIAcuity Software Suite 1.1.3 that you would like to import and use in QIAcuity Software Suite 1.2, you will need to import the plates before upgrading from version 1.1.3 to version 1.2. You may then export the plates again. Future software version starting from QIAcuity Software Suite 2.0 will facilitate import of plates from previous QIAcuity Software Suite versions.
The new improvements are as follows:
-Support for the Nanoplate 8.5k 24-well
-Hyperwell functionality to combine several wells to one combined well for analysis
-Automated plate archiving functionality
-Functionality to show the number of single/double positives in 2D scatterplots
-VPF (Volume Precision Factor) to further improve concentration calculation (see related resources)
-Additional improvements for stabilization and troubleshooting
Version 2.2
The QIAcuity Control Software is an integral part of the QIAcuity instrument. It offers a GUI (graphical user interface) for basic functionalities such as plate setup, changing the order of plates to be processed, and monitoring the status of runs in real time. After a run is completed, the data are stored on the instrument’s memory and are sent to the connected QIAcuity Software Suite for analysis.
The new Version 2.2 of the QIAcuity software (QIAcuity Software Suite v 2.2 and QIAcuity CSW v 2.2.) offers improvements for users working under GMP by adding a user ID validation during the report signing and the addition of timezone offset stamp for audit trail entries and for result report data. Furthermore, the addition of a standard deviation and coefficient of variance calculation in percentage of mean concentration calculation for replicates where implemented. In addition, the instrument camera stability was improved and an internal validation method was implemented.
Detailed information about the QIAcuity Control Software version 2.2 is available in the Release Note, which can also be downloaded under section “Software Release Notes”.
Note: The latest CSW version 2.2 is only compatible with the Software Suite version 2.2. If only one software component is updated, no connection between the Software Suite and the CSW can be established.
Important: Please follow the instructions provided in the QIAcuity User Manual for software version 2.2 and in the Release Note.
It is strongly recommended to update the QIAcuity Software Suite first before proceeding with the QIAcuity Control Software.
Note: After clicking reboot during CSW upgrade or change of Suite connection, login screen may appear for short period. Please ignore it and wait for the QIAcuity instrument to shut down and restart itself.
Please contact QIAGEN Technical Services if you are unsure and require technical support.
SHA1 checksum: 3892D7A434A8F072A15008C76EB088BB78F1C255
July 2021
FAQ
The plate is designed for a single use run. For example, even if only 30 samples are loaded into the 96-well plate, a whole plate will be sealed by the roller. It can't be unsealed and used for another run. The QIAcuity Software won’t allow to set up a separate experiment for the same nanoplate to avoid that previously processed plates being not partitioned a second time.
The QIAcuity EG PCR Kit should be stored immediately upon receipt at –30 to –15°C in a constant-temperature freezer and protected from light. The QIAcuity EG PCR master mix can also be stored protected from light at 2–8°C. Components are stable for 6 months, unless otherwise indicated on the label.
The QIAcuity Nanoplates does not have expiry date and are stable for at least 1 year when stored at RT.
A standard PCR plate is required to set up dPCR reaction before transferring it to the nanoplate to ensure a proper mixing of the reaction mix before partitioning.
QIAGEN dPCR assays (such as dPCR LNA Mutation Assays) can be found on https://geneglobe.qiagen.com/.
This is not needed. The QIAcuity is equipped with a flexible power supply technology and operates within a range of 100–240V AC, 50/60 Hz, 1500 VA (max).
If you had run a nanoplate for which the installed VPF misses the specific factor, the software will notify you. If you then analyze without the specific VPF, the impact depends on the variation of the partition volume of the new Nanoplate batch compared to the latest. Typically this variation is ±6–7% (approx. 5% CV over the entire plate). The analysis may be repeated after updating the VPF file. After installing the latest VPF and re-analysis of the run, a copy of the plate is generated in the QIAcuity Software Suite including the new results.
QIAGEN master mixes are optimized for nanoplate microfluidics and are recommended to be used with our dPCR system. They also include an optimized reference dye required for proper analysis.
An essential temperature gradient functionality was introduced with software version 2.5. When updating older software versions to 2.5, each QIAcuity instrument will offer the temperature gradient and may be used to find the best cycling temperature for new dPCR assays. When running a QIAcuity Four or a QIAcuity Eight, all plates may have their own temperature profile, including the option for a temperature gradient.
QIAGEN offers a complete range of nucleic acid purification systems. These include QIAprep kits for purification of plasmid DNA, QIAamp, and DNeasy kits for purification of genomic DNA, RNeasy kits for purification of total RNA, and the PAXgene Blood RNA System for stabilization and purification of RNA from blood. Phenol and other contaminants can be efficiently removed from crude RNA preps using the RNeasy MinElute Cleanup Kit to clean up and concentrate RNA for sensitive assays. Details about QIAGEN kits for nucleic acid purification can be found at www.qiagen.com.
If you had analyzed your nanoplates without VFP, the impact depends on the variation of the partition volume of the new nanoplate batch compared to the latest. The VPF reduces the CV from approximately 5% to 2%. We recommend to reanalyze results in case the data originated from different wells (e.g., copy number variations or gene expression data sets for which the reference sample was measured in a different well). Results obtained across different plates should also be r-analyzed. A reanalysis is not required for assay data that were analyzed within the same well (e.g., mutation rate determination using two channels within the same well).
In dPCR we measure the absolute concentration of targets at endpoint reaction. Concentrations of unknowns can be determined based on dPCR results observed (number of negatives, number of positives, and partition volume analyzed).
In general, nanoplates provide partitions of fixed sizes that enable a very precise way of sample concentration calculation. If a new molding form is used for nanoplate manufacturing, potential variation of partition sizes can be addressed by applying the molding form-specific VPF. Thus, each time a new molding form is used, a new VPF is created and made available. Currently, the VPF is updated once every 3–6 months.
Nanoplate 8.5K 24-well:12 μl
Nanoplate 8.5K 96-well: 12 μl
The fluorescent signal in the reference channel is measured to determine the number of valid partitions in a well. In addition, differences in the signal intensities between partitions are normalized and the fluorescent signals in the target channels are corrected accordingly.
The VPF provides a set of well-specific and molding form-specific factors used to specify the exact reaction volume of a nanoplate, thus increasing the concentration calculation of each well.
Yes, the report includes a notification if the matching VPF was missing and, therefore, not applied to the analysis. If the matching VPF was applied there is no notification on the report.
The QIAcuity Probe PCR Kit should be stored immediately upon receipt at –30 to –15°C in a constant-temperature freezer and protected from light. The QIAcuity Probe PCR master mix can also be stored protected from light at 2–8°C. Components are stable for 12 months, unless otherwise indicated on the label.
The QIAcuity EG PCR Kit should be stored immediately upon receipt at –30 to –15°C in a constant-temperature freezer and protected from light. The QIAcuity EG PCR master mix can also be stored protected from light at 2–8°C. Components are stable for 6 months, unless otherwise indicated on the label.
The QIAcuity Nanoplates does not have expiry date and are stable for at least 1 year when stored at RT.
Nanoplate 26K 24-well: 40 μl
Nanoplate 8.5K 24-well:12 μl
Nanoplate 8.5K 96-well: 12 μl
All DNA samples used in reaction mixes should show similar quality and quantity, which can easily be assessed using UV spectrophotometry. DNA samples with an average length of ≥20 kb (e.g., genomic DNA purified via spin column with silica membrane) should be fragmented by restriction digestion before partitioning. Enzymatic fragmentation of larger DNA ensures an even distribution of template throughout the QIAcuity Nanoplate, which in turn leads to an accurate and precise quantification.
The instrument software GUI shows error codes including a description and information how to resolve the error. The instrument touchscreen shows an alarm icon in the upper right corner that turns red in case of an instrument failure. Accessing the System Status in the Tool tab allows users to clear errors. Rebooting of the instrument is required to complete the removal of the error. Please do not skip this step. You may always contact QIAGEN Technical Services in case of any question.
The QIAcuity instrument software does not allow to read and process a plate without seal. If you would like to perform a dry run please use sealed plate and set up this plate in the QIAcuity Software Suite.
dPCR master mix can be used in qPCR to optimize sample concentration and/or primer/probe concentration prior to assay transfer from qPCR to dPCR.
The user manual contains instructions on how to perform a regular cleaning and decontamination, and how to replace air filters on the QIAcuity instruments. A regular maintenance reduces the dust in the instrument and therefore minimizes the presence of dust particles on the nanoplate, which might interfere with the plate analysis.
The up-to-date VPF can be downloaded in the resource section of the QIAcuity product webpage. The VPF is compatible with the QIAcuity Software Suite 1.2 and higher. The latest VPF file contains all factors for all existing nanoplate batches. If a nanoplate from a new nanoplate batch is run and the latest version of the VPF was not installed, the software will recognize this and will give a message to install the latest VPF file. Please note that Applying the VPF file cannot be reversed.
The QIAcuity reads emitted fluorescence from the bottom of the plate, which is covered with a foil. For best results, keep the foil clean and avoid damages such as scratches. Also, keep the barcode on the side of the plate clean and intact. Ensure that you wear gloves when working with a plate and do not apply force to it. For a safe handling of the plate, please place the plate into a nanoplate tray.
No. The QIAcuity platform introduces four variations: QIAcuity One 2plex, QIAcuity One 5plex, QIAcuity Four (5plex), and QIAcuity Eight (5plex). All of them have fix channel combination.
Results are stored as part of the plates within the QIAcuity Software Suite. In version 1.1.2, plates can be exported to another file location, for example, an external HDD, and imported again if needed. From version 1.2 onwards, plates can also be archived automatically. To do so, an archive destination has to be defined. Additional information can be found in the user manual.
Both software are designed to be upgraded by users. The user manual includes instructions on how to perform the upgrades; instructions for the QIAcuity Suite Software upgrade can be found at page 38 and instructions for the CSW upgrade on page 67 (QIAcuity User Manual, 03/2021).
