LoopLigase (50 U)
Cat no. / ID. L6130L
Features
- Faster reaction rate
- Minimal template bias
- Circularizes longer substrates
- Does not require splint oligonucleotides
- Does not generate concatemers
Product Details
LoopLigase is a thermostable single-stranded ligase capable of circularizing ssDNA or ssRNA templates. It catalyzes the formation of a phosphodiester bond between the terminal 5'-monophosphate and 3'-hydroxyl groups of an ssDNA or ssRNA. It is a thermostable, ATP-dependent enzyme for direct ligation without relying on adjacent complementary regions.
The enzyme is supplied with 5x LoopLigase buffer.
Performance
LoopLigase shows high-efficiency circularization of both ssDNA and ssRNA (see “ Exceptionally efficient intramolecular ligation”).
LoopLigase circularizes longer substrates (see “ Efficient circularization of longer substrates”) and does not require a splint oligonucleotide while generating low to no circular concatemers (see “ No splint oligonucleotide required and no concatemers generated”). LoopLigase shows minimal template bias (see “ Minimal bias for terminal nucleotides”).
Properties:
Storage temperature: –25°C to –15°C
Molecular weight: 44.05 kDa
| Test | Specifications |
|---|---|
| SDS purity | >99% |
| Single-stranded exonuclease | <1.0% Released |
| Double-stranded exonuclease | <1.0% Released |
| Double-stranded endonuclease | No conversion |
| E. coli DNA contamination | <10 copies |
| Non-specific RNase | No detectable non-specific RNase |
Principle
Source of Protein: A recombinant E. coli strain carrying the cloned LoopLigase gene.
Unit Definition: One unit is the amount of LoopLigase required to ligate 50% of 0.2 µM of 64 nt linear ssDNA fragments (with 5’-monophosphate and 3’-hydroxyl groups) into circular ssDNA in 15 minutes at 60°C in a 20 μL reaction with 1x LoopLigase reaction buffer.
Procedure
Usage Instructions: Circularization of ssDNA with 5’- monophosphate and 3’-hydroxyl groups
1. Set up the following reaction mixture in a total volume of 20 µL:
| Components | Final Concentration | Volume |
| Nuclease free water | N/A | 14 µL |
| 5x LoopLigase Buffer (B6130) | 1x | 4 µL |
| ssDNA substrate | 0.2 pmol/µL | 1 µL |
| LoopLigase (L6130) | 2 U | 1 µL |
| Total volume = | 20 µL |
2. Incubate at 60°C for 15–60 minutes.
3. Reaction can be stopped by incubation at 85°C for 10 minutes.
Notes:
5x LoopLigase Buffer: It is recommended that the reaction buffer be discarded after one year of storage at –20°C and replaced with fresh buffer to ensure maximum performance.
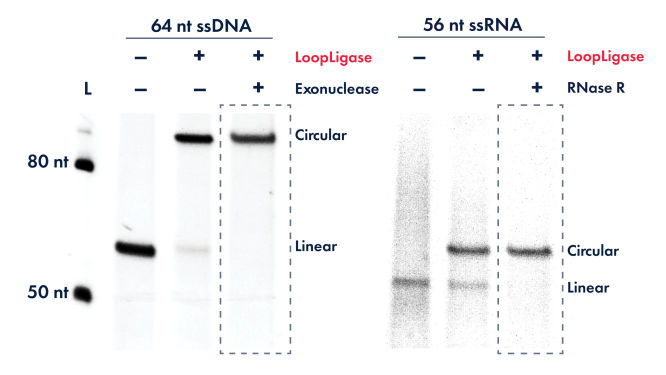
Removing unligated linear substrates: After completion of the circularization reaction, any remaining unligated linear DNA can be removed using exonuclease digestion. To ensure digestion of any possible secondary structures (e.g., hairpins), a combination of Exonuclease I (cat. no. X8010L, ssDNA specific) and Exonuclease III (cat. no. X8020L, dsDNA specific) can be used. RNase R can be used to remove linear RNA from the circularization reaction. Circular products of LoopLigase are resistant to exonucleases.
Quality Control Analysis:
Protein concentration (OD280) is determined by OD280 absorbance.
Physical purity is evaluated by SDS-PAGE of concentrated and diluted enzyme solutions followed by silver stain detection. Purity is assessed by comparing the aggregate mass of contaminant bands in the concentrated sample to the mass of the protein of interest band in the diluted sample.
Single-stranded exonuclease is determined in a 50 µL reaction containing a radiolabeled single-stranded DNA substrate and 10 µL of enzyme solution incubated for 4 hours at 37°C.
Double-stranded exonuclease is determined in a 50 µL reaction containing a radiolabeled double-stranded DNA substrate and 10 µL of enzyme solution incubated for 4 hours at 37°C.
Double-stranded endonuclease is determined in a 50 µL reaction containing 0.5 µg of plasmid DNA and 10 µL of enzyme solution incubated for 4 hours at 37°C.
E. coli 16S rRNA contamination is evaluated using 5 µL replicate samples of enzyme solution denatured and screened in a TaqMan qPCR assay for the presence of contaminating E. coli genomic DNA using oligonucleotide primers corresponding to the 16S rRNA locus.
Non-specific RNase contamination is assessed using the RNase Alert kit (Integrated DNA Technologies), following the manufacturer’s guidelines.
Applications
LoopLigase is used for the following applications:
- Template circularization for rolling circle amplification (RCA)
- Preparation of circular libraries for NGS
Supporting data and figures
Exceptionally efficient intramolecular ligation
LoopLigase shows high-efficiency circularization of both ssDNA and ssRNA. Unligated ssDNA and ssRNA were digested by exonuclease and RNase R, respectively.