✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Cat. No. / ID: 206544
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Reliable and accurate detection of subtle sequence variations
- Highly suited for use with any cycler with HRM capabilities
- Distinct melting curves due to novel EvaGreen fluorescent dye
- Easy development of reliable new HRM genotyping assays
- Convenient master mix format and optimized protocols
Product Details
The Type-it HRM PCR Kit provides a reliable solution for fast and accurate genotyping via HRM analysis. The kit is optimized to even enable successful analysis of genomic loci that are difficult to amplify. The kit is highly suitable for mutation scanning and can be used to screen samples for unknown mutations. Development of new HRM genotyping assays is easy and straightforward. Specific amplification products, reduced nonspecific amplification, and reliable results are consistently ensured — time-consuming optimization of PCR parameters is no longer required. This leads to standardization and flexibility, in addition to substantial time and cost savings. The kit is compatible with all real-time instruments suitable for HRM analysis, such as the Rotor-Gene Q, the Rotor-Gene 6000, the LightCycler 480, and the Applied Biosystems 7500 Fast System.
Performance
The Type-it HRM PCR Kit is validated for accurate resolution of sequence variations and is highly suitable for unambiguous allelic discrimination using innovative HRM technology. With HRM technology, previously unknown and even complex sequence variations, as seen in challenging genotyping applications, can be readily detected and characterized in an easy and straightforward way (see figure " Highly accurate genotyping").
Reliable and accurate detection of subtle sequence variations
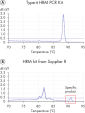
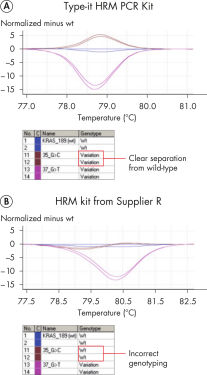
The master mix provided with the kit contains the novel double-stranded DNA-binding fluorescent dye, EvaGreen, and includes an optimized HRM buffer, HotStarTaq Plus DNA Polymerase, Q-Solution, and dNTPs. Together, these components ensure PCR specificity and provide reliable results, even with difficult genomic loci. The Type-it HRM PCR Kit outperformed kits tested from other suppliers and enables amplification of even the most challenging subtle sequence differences. The unique composition of the Type-it HRM PCR Buffer ensures highly stringent and targeted primer binding compared to other commercially available HRM master mix chemistries (see figure " Highly specific and successful amplification of difficult genomic loci"). In contrast to kits from other suppliers, the Type-it HRM PCR Kit delivers consistent performance at the first attempt on all real-time instruments tested, ensuring highly accurate detection of even Class IV SNPs (see figure " Successful genotyping of an A/T Class IV SNP") and gene mutations (see figure " Successful typing of gene mutations"). These advantages also result in significant time and cost savings by minimizing the number of samples that need to be retested.
Fast and easy mutation scanning
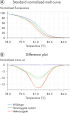
Highly reliable mutation scanning can be performed with the Type-it HRM PCR Kit. Unknown gene mutations (insertions and deletions) are readily detected and reliably distinguished, resulting in distinct melting curves (see figure " Successful mutation scanning").
See figures
Principle
HRM is a closed-tube, post-PCR analysis method that has raised enormous scientific interest. HRM characterizes double-stranded PCR products based on their dissociation (melting) behavior as they transition from double-stranded DNA (dsDNA) to single-stranded DNA (ssDNA) with increasing temperature. PCR products can be discriminated according to sequence, length, GC content, or strand complementarity, down to single base pair changes. Previously unknown and even complex sequence variations as seen in challenging genotyping applications can be easily analyzed (see figure " Highly accurate genotyping"). HRM is easier and more cost-effective than probe-based genotyping analysis and, unlike conventional methods, it prevents carry-over contamination of PCR products.
The unique kit components ensure highly specific amplification (see table).
Novel EvaGreen dye for distinct melting curves
The Type-it HRM PCR Kit contains EvaGreen, a third-generation, saturating fluorescent dye which selectively binds to double-stranded DNA. In contrast to conventional SYBR® Green I, EvaGreen can be used at higher concentrations without PCR inhibition and shows equal binding affinity for GC-rich and AT-rich regions with no apparent sequence preference. This makes EvaGreen highly suitable for HRM analysis of all types of PCR products, enabling distinct melting curves due to visualization of lower fluorescent differences, thereby ensuring standardized results.
Optimized HRM PCR Master Mix
HRM PCR Master Mix consists of HotStarTaq Plus DNA Polymerase and an innovative HRM PCR buffer system, both of which enable highly specific amplification and distinct melting curves of even difficult genomic loci such as Class IV SNPs (see figure " Successful genotyping of an A/T Class IV SNP"). Specific amplification of the desired PCR product is ensured and generation of nonspecific products and formation of primer–dimers is minimized, if not eliminated (see figure " Successful typing of gene mutations").
The innovative buffer (included in the master mix) maintains specific amplification in every cycle of PCR by promoting a high ratio of specific-to-nonspecific primer binding during the annealing step in each PCR cycle. Owing to a uniquely balanced combination of KCl and (NH4)2SO4, the buffer provides stringent primer-annealing conditions over a wider range of annealing temperatures and Mg2+ concentrations than conventional PCR buffers. Optimization of PCR by varying the annealing temperature or the Mg2+ concentration is therefore often minimal or not required.
Suitable for difficult mutation loci
The Type-it HRM PCR Kit is a powerful system for easy detection of difficult mutation loci (see figure " Highly specific and successful amplification of difficult genomic loci"). Mutations or SNPs located in GC-rich regions or regions of high secondary structure are difficult to amplify using conventional HRM PCR solutions. The 2x HRM PCR Master Mix includes the innovative PCR additive, Q-Solution, at a defined concentration to help overcome these challenges. Q-Solution improves PCR by modifying the melting behavior of DNA, resulting in successful amplification even of difficult target sequences without the need for optimization.
| Contents | Benefits |
|---|---|
| 2x HRM PCR Master Mix | Eliminates the need for optimization in the development of new assays Specially developed for HRM analysis of mutations and SNPs Dedicated for use with all cyclers suitable for HRM analysis Convenient reaction setup, minimizing pipetting errors |
| HotStarTaq Plus DNA Polymerase | Highly specific amplification Fast and easy room-temperature reaction setup |
| Type-it HRM PCR Buffer | Distinct melting curves Increased amplification specificity |
| Q-Solution | Improves amplification of difficult templates |
| EvaGreen Dye | Novel, saturating dsDNA-binding dye Distinct melting curve analysis |
See figures
Procedure
The Type-it HRM PCR Kit includes dedicated, application-specific protocols, preoptimized for use with various real-time cyclers such as the Rotor-Gene Q, Rotor-Gene 6000, and LightCycler 480. In addition, the Applied Biosystems 7500 Fast System and Applied Biosystems 7900 can also be used with the optimized protocols provided with the kit. A detailed protocol is available on our Web site.
Fast-cycling procedure and easy-to-develop HRM assays for straightforward results
The unique features of the Type-it HRM PCR Buffer and optimized protocols provided with the kit ensure successful HRM genotyping results at the first attempt. Lengthy optimization of reaction parameters is not required and new HRM genotyping assays can be quickly and easily standardized and implemented into routine research (see figure " Successful HRM analysis without the need for optimization"). In addition, the fast cycling protocol of the Type-it HRM PCR Kit increases throughput by shortening PCR run times, enabling accurate results to be achieved faster.
Convenient kit format
The kit is provided in a ready-to-use, preoptimized master mix format for greater convenience. Use of a master mix saves time, simplifies handling for reaction setup, and increases reproducibility by eliminating many possible sources of pipetting errors and contamination — pipetting steps are minimized and tedious calculations are eliminated. Room-temperature reaction setup using the master mix is fast and easy. HotStarTaq Plus DNA Polymerase (included in the master mix) is activated by a 5-minute, 95°C incubation step, which can easily be incorporated into existing thermal cycling programs.
See figures
Applications
The Type-it HRM PCR Kit can be used for several applications such as:
- Detection of deletions, insertions, and translocations
- Scanning for gene mutations
- SNP genotyping
- Detection and discrimination of microbial variants
The Type-it HRM PCR Kit is a universal tool applicable in a range of research fields, including:
- Typing of disease and cancer loci
- Biomarker discovery
- Disease association studies
- Typing of transgenic plants and animals
- Pathogen detection and genotyping
Supporting data and figures
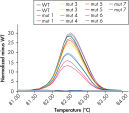
Successful typing of gene mutations.
Specifications
| Features | Specifications |
|---|---|
| Applications | Detection of SNPs, mutations and mutation scanning |
| Product use | Functionally validated and developed for reliable detection of genetic differences using HRM |
| Reaction type | PCR amplification |
| Enzyme activity | 5'-> 3' Exonuclease activity |
| With or without ROX | Without ROX |
| Sequence specific Probe | Not necessary, EvaGreen dye for detection included in the Mastermix |
| Sample/target type | Genomic DNA |
| With/without hotstart | With |
| Mastermix | Yes |
| Real-time or endpoint | Both |