✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Cat. No. / ID: 80234
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Maximum output with minimal sample consumption
- Releases DNA/RNA without compromising integrity
- Effectively separates RNA and DNA
- Comprehensive DNA and RNA analysis of the same FFPE sample
Product Details
For reliable comparison of genomic and transcriptomic data, purification of DNA and RNA from the same sample is essential. The AllPrep DNA/RNA FFPE Kit uses a patent-pending solubilization method to purify DNA and RNA from formalin-fixed, paraffin-embedded (FFPE) tissue sections. Purified analytes are suitable for use in applications such as real-time PCR and Pyrosequencing. The kit can be automated on the QIAcube Connect.
Performance
See figures
Principle
The AllPrep DNA/RNA FFPE Kit is specially designed for simultaneous purification of genomic DNA and total RNA from FFPE tissue sections. Pure DNA and RNA are obtained from the entire sample, in contrast to other procedures where the biological sample is divided into two before being processed separately. Simply dividing a sample in half for separate DNA and RNA purification procedures results in the purification of DNA and RNA from different populations of cells, which may differ in their properties. Purification of DNA and RNA from the same sample also helps to prevent waste, since FFPE samples are precious, often difficult to retrieve, and limited in amount.
Due to fixation and embedding conditions, nucleic acids in FFPE samples are usually heavily fragmented and are often of a lower molecular weight than those obtained from fresh or frozen samples. A major obstacle to isolating DNA and RNA from the same FFPE sample is that fragmented DNA is short and can be partly single-stranded and therefore more closely resembles RNA than intact DNA. This property of fragmented DNA makes physical separation of DNA and RNA difficult. The AllPrep DNA/RNA FFPE Kit uses a patent-pending solubilization method to differentially release DNA and RNA from a single FFPE sample.
Nucleic acids in FFPE samples are also chemically modified by formaldehyde. Although formaldehyde modification cannot be detected in standard quality control assays, such as gel electrophoresis or lab-on-a-chip analysis, it does strongly interfere with enzymatic analyses. The AllPrep DNA/RNA FFPE Kit is optimized to reverse as much formaldehyde modification as possible without further DNA and RNA degradation.
Procedure
See figures
Applications
The AllPrep DNA/RNA FFPE Kit is optimized to reverse as much formaldehyde modification as possible without further DNA and RNA degradation; however nucleic acids purified from FFPE samples should not be used in downstream applications that require high-molecular-weight DNA or full-length RNA. Some applications may require modifications to allow the use of fragmented nucleic acids (e.g., designing small amplicons for PCR and RT-PCR). For cDNA synthesis, gene-specific primers should be used instead of oligo-dT primers. If it is not possible to use gene-specific primers, random primers should be used.
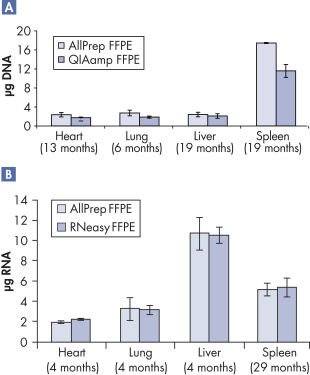
Supporting data and figures
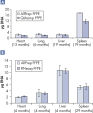
Purification of DNA and RNA from FFPE samples.
Specifications
| Features | Specifications |
|---|---|
| Applications | PCR, qPCR, real-time RT-PCR, microarray |
| Elution volume | RNA: 14-30µl ; DNA: 30-100µl |
| Purification of total RNA, miRNA, poly A+ mRNA, DNA or protein | RNA (miRNA) and DNA |
| Processing | Manual (centrifugation) |
| Format | Spin column |
| Number of preps per run | 50 |
| Sample amount | max. 4*10 µm sections or 2*20 µm sections |
| Technology | Silica technology |
| Time per run or per prep | 6h for 10 samples, including sectioning |
| Yield | Varies |
| Main sample type | FFPE tissue samples |