Cat. No. / ID: 9001861
Features
- Outstanding thermal and optical performance due to rotary format
- An unmatched optical range spanning UV to infrared wavelengths
- State-of-the art analyses supported by user-friendly software
- Low maintenance and maximum convenience due to robust design
- High performance in multiple applications with QIAGEN assays
Product Details
Explore the virtual demo to learn more about the Rotor-Gene Q.
Note that the Rotor-Gene Q instruments are a life science product and not intended for in vitro diagnostic use.
Performance
See figures
Principle
Unique rotary design for outstanding performance
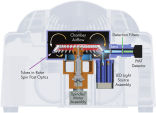
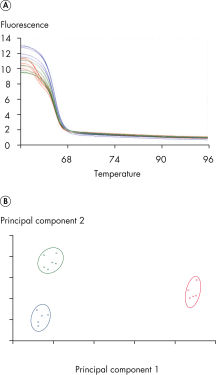
The unique centrifugal rotary design of the Rotor-Gene Q makes it the most precise and versatile real-time PCR cycler currently available (see figure " Cross-section of the Rotor-Gene Q"). Each tube spins in a chamber of moving air, keeping all samples at precisely the same temperature during rapid thermal cycling. Detection is similarly uniform. When each tube aligns with the detection optics, the sample is illuminated and the fluorescent signal is rapidly collected from a single, short optical pathway. This thermal and optical uniformity results in sensitive, precise, and fast real-time PCR analysis (see figure " Precise real-time PCR analysis"). It also eliminates sample-to-sample variations and edge effects. These are unavoidable in traditional block-based instruments due to temperature gradients across the block and multiple, complex optical pathways.
The rotary design delivers:
- Tube-to-tube variation ±0.02°C
- Uniform detection eliminating the need for ROX reference dye
- Fast ramping and negligible equilibration times for short run-times
- Complete confidence in your results
Unrivaled optical range enables multiple applications
Whether your assay is based on intercalating dyes such as SYBR® Green, probes such as hydrolysis (TaqMan), hybridization (FRET), Scorpion probes, or other multiplex chemistries, the Rotor-Gene Q meets your requirements. With up to 6 channels spanning UV to infrared wavelengths, the cycler delivers the widest optical range currently available (see table). In addition, the software allows you to create new excitation/detection wavelength combinations, which means that the Rotor-Gene Q is compatible with dyes you may use in the future.
| Channel | Excitation (nm) | Detection (nm) | Examples of fluorophores detected |
|---|---|---|---|
| Blue | 365 ± 20 | 460 ± 20 | Marina Blue, Edans, Bothell Blue, Alexa Fluor 350, AMCA-X |
| Green | 470 ± 10 | 510 ± 5 | FAM, SYBR® Green I, Fluorescein, EvaGreen, Alexa Fluor 488 |
| Yellow | 530 ± 5 | 557 ± 5 | JOE, VIC, HEX, TET, MAX, CAL Fluor Gold 540, Yakima Yellow |
| Orange | 585 ± 5 | 610 ± 5 | ROX, CAL Fluor Red 610, Cy 3.5, Texas Red, Alexa Fluor 568 |
| Red | 625 ± 5 | 660 ± 10 | Cy5, Quasar 670, LightCycler Red640, Alexa Fluor 633 |
| Crimson | 680 ± 5 | 712 high pass | Quasar 705, LightCycler Red705, Alexa Fluor 680 |
| HRM | 460 ± 20 | 510 ± 5 | SYBR® Green I, SYTO9, LC Green, LC Green Plus+, EvaGreen |
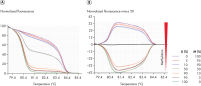
Expand your research with HRM
High-resolution melting analysis (HRM) is a closed-tube, post-PCR analysis that has raised enormous scientific interest. HRM characterizes double-stranded PCR products based on their dissociation (melting) behavior. It is similar to classical melting curve analysis, but provides far more information for a wider range of applications. PCR products can be discriminated according to sequence, length, GC content, or strand complementarity, down to single base-pair changes. Previously unknown and even complex sequence variations can be readily detected and characterized in a robust and straightforward way. The rotary design of the Rotor-Gene Q and its outstanding thermal and optical performance are highly suited to HRM.
The HRM option for the Rotor-Gene Q includes:
- A specially tuned high-intensity optical HRM channel
- Thermal resolution of 0.02°C
- High data-acquisition rates
- Comprehensive HRM software
Minimum maintenance; maximum convenience
The Rotor-Gene Q is engineered to reduce the need for maintenance and to maximize ease of use. This saves time and costs, and allows you to focus on your research; not on keeping the cycler up and running.
Convenient features of the Rotor-Gene Q include:
- Lifetime guarantee on highly stable LEDs, no expensive lamps or lasers to change, no gradual performance loss of light source
- No optical calibration needed at installation or when the instrument is moved
- No sample block to clean
- No condensation or bubbles in reactions due to rotation
- Small, light, and robust; simply place the instrument wherever you like
See figures
Procedure
Flexible formats match your workflows
The Rotor-Gene Q supports multiple PCR tube formats to suit a range of needs (see figure " Flexible formats"). Changing the format, by simply switching the snap-fit metal rotor that holds the tubes, takes just seconds. As well as tubes, Rotor-Discs are available, which offer accelerated setup and higher throughput. Rotor-Discs are circular plates of vertically oriented reaction wells. The Rotor-Disc 100 is the equivalent of a 96-well plate with an additional 4 reference wells. These extra wells can be conveniently used for more reactions or additional controls. Alternatively, the Rotor-Disc 72 has 72 wells. Rotor-Discs can be quickly and easily sealed with plastic film using a Rotor-Disc Heat Sealer. For all you need to run reactions using Rotor-Discs, choose the Rotor-Disc 100 Starter Kit or the Rotor-Disc 72 Starter Kit.
You can perform manual reaction setup, or take advantage of QIAGEN's automated solutions for reaction setup. The QIAgility is cost-effective and delivers rapid, high-precision PCR setup, while the QIAsymphony AS is ideal for laboratories performing routine PCR tests on a day-to-day basis. Both instruments perform automated reaction setup in Rotor-Gene formats, allow direct transfer of sample lists, and are supplied with verified protocols for real-time PCR master mixes.
Easy routine verification
Laboratories may often want to verify thermal accuracy. For most cyclers, this requires interaction with a service engineer. With the Rotor-Gene Q, this is not necessary. Instead, the easy-to-use, cost-effective Rotor-Disc OTV (Optical Temperature Verification) Kit automates accuracy testing. The full procedure takes only a couple of minutes.
See figures
Applications
A range of QIAGEN kits for the Rotor-Gene Q enables reliable quantification in all your real-time PCR applications without the need for optimization of reaction and cycling conditions. Kits for real-time PCR and HRM applications are available for:
- Gene expression analysis
- Pathogen detection
- DNA methylation analysis
- Genotyping and gene scanning
- miRNA research
Software
Q-Rex Software
Q-Rex Software is a new operating and analysis software for the Rotor-Gene Q, providing several unique new features that promote a more user-friendly interface to help streamline your qPCR workflow. The software is suitable for use by the most novice researchers, while maintaining the highly complex data analysis functions required by advanced researchers.
Rotor-Gene Q Software
The comprehensive Rotor-Gene Q software package supports all current state-of-the art real-time analysis procedures from basic to advanced algorithms. This provides complete freedom to analyze your valuable experimental data and increases the reliability of your results. Data security is assured and all process steps are trackable from starting the run to exporting the results.
Superior software available for genotyping and mutation detection using HRM analysis
Rotor-Gene ScreenClust HRM Software is an extension to the Rotor-Gene operating software. This software is the most powerful tool currently available for analysis of HRM data from the Rotor-Gene Q or Rotor-Gene 6000 cycler. By grouping samples into clusters, Rotor-Gene ScreenClust HRM Software opens a new dimension in HRM analysis for applications such as genotyping and mutation screening.
REST software 2009
REST software 2009 is a standalone software tool for analysis of gene expression data from quantitative real-time PCR experiments. REST software 2009 is available for download under the "Resources" tab, and provides valuable analysis, including:
- Estimation of up- and down-regulation for gene expression studies
- Randomization and bootstrapping techniques
- Graphical data output via whisker-box plots
Traditional relative quantification enables estimation of gene expression. However, this method does not provide statistical information that is suitable for comparing expression in groups of treated and untreated samples in a robust manner. The integrated randomization and bootstrapping methods used in REST software 2009 test the statistical significance of calculated expression ratios and can be used even when the data includes outliers.
REST software 2009 applies a mathematic model that takes into account the different PCR efficiencies of the gene of interest and reference genes. Compared to using a single reference gene, using multiple reference genes for normalization can improve the reliability of results.
Supporting data and figures
Identification of a class IV SNP.
Specifications
| Features | Specifications |
|---|---|
| Protocols/main application on this instrument | Gene expression analysis, microRNA detection, Virus detection, SNP genotyping, SNP genotyping, High Resolution Melt analysis (HRM) |
| Heat dissipation/thermal load | Average, 0.183 kW (632 BTU/hour); Peak, 0.458 kW (1578 BTU/hour) |
| Weight | 12.5 kg (27.6 lb.), standard configuration |
| Features | Dynamic range, 10 orders of magnitude |
| Transportation conditions | –25ºC to 60ºC (–13ºF to 140ºF) in manufacturer’s package; Max. 75% relative humidity (noncondensing); Environmental class 2K2 (IEC 60721-3-2) |
| Overvoltage category | II |
| Warranty | 1 year on instrument; lifetime guarantee on excitation LEDs |
| Operating temperature | 18-30ºC (64-86ºF) |
| Place of operation | For indoor use only |
| Storage conditions | 15ºC to 30ºC (59ºF to 86ºF) in manufacturer’s package; Max. 75% relative humidity (noncondensing); Environmental class 1K2 (IEC 60721-3-1) |
| Samples per run (throughput) | Tubes 0.2 ml; Strip Tubes 0.1 ml (4 tubes); Rotor-Disc 72; Rotor-Disc 100; Up to 100 samples per run using a Rotor-Disc 100 |
| Typical run time | 40 cycles in 45 min with the QIAGEN RG Kits (assay dependent) |
| Dimensions | Width, 37 cm (14.6 in.); Height, 28.6 cm (11.3 in.); Depth (without cables), 42 cm (16.5 in.); Depth (door open), 53.8 cm (21.2 in.) |
| Power | 100–240 V AC, 50–60 Hz, <520 VA (peak); Power consumption <60 VA (standby); Mains supply voltage fluctuations are not to exceed 10% of the nominal supply voltages; F5a 250 V fuse |
| Kits designed for this instrument | artus QS-RGQ Kits (not available in all countries), RG SYBR Green PCR Kits; RG SYBR Green RT-PCR Kit; RG Probe PCR Kits; RG Probe RT-PCR Kit; RG Multiplex PCR Kit |
| Humidity | 10–75% (noncondensing) |
| Thermal performance | Temperature range, 35 to 99ºC; Temperature accuracy, ±0.5ºC (type, measured 30 seconds after clock start); Temperature resolution, ±0.02ºC (smallest programmable increment); Temperature uniformity, ±0.02ºC; Ramp rate (peak ramp rates, air), >15ºC/s heating, >20ºC/s cooling |
| Pollution level | 2; Environmental class 3K2 (IEC 60721-3-3) |
| Software | Rotor-Gene Q software, supplied on the installation CD provided |
| Optical System | Up to 6 channels spanning UV to infra-red wavelengths; Excitation sources: High energy light-emitting diodes; Detector: Photomultiplier; Acquisition time: 4 s. The software allows to create new excitation/detection wavelength combinations |
| Technology | Real-time PCR cycler |
| Altitude | Up to 2000 m (6500 ft) |