Products

QuantiNova LNA Probe PCR Custom Panels
Cat no. / ID. 249975
Features
- Fully customizable panel containing any of our 1.3 million predesigned assays and/or custom assays
- Easy online ordering through the intuitive custom builder tool
- Exceptional sensitivity and specificity using short, LNA-enhanced primers and FAM-labeled hydrolysis probes
- Stability for room-temperature setup allowing automation
- Robust and reproducible results in less than 2 hours using the QuantiNova Probe PCR Kit
Product Details
QuantiNova LNA Probe PCR Custom Panels enable quick and reliable gene expression profiling using hydrolysis probe-based qPCR. These fully customizable panels allow you to choose from any of our predesigned QuantiNova LNA Probe PCR Assays and/or your own custom assays. LNA enhancement enables us to design shorter primers with optimal Tm, allowing flexible positioning on the target for optimal sequence discrimination. Thorough design validation ensures top performance and robust detection. Optimized using QuantiNova chemistry, the simple workflow can be completed in under 2 hours and is robust enough to support room-temperature setup, automation and delayed run. We also offer Flexible Panels, which are semi-customizable from the assays in our available Focus Panels.
Need a quote for your research project or would you like to discuss your project with our specialist team? Just contact us!
Performance
Expert LNA enhancement for top qPCR performance
All QuantiNova LNA Probe PCR Assays are developed using stringent design criteria and lab-validated algorithms. Twenty years of LNA design experience has enabled us to optimize our sophisticated LNA design algorithm, which incorporates over 50 different parameters to guarantee the most optimal assay for successful target detection. Each primer–probe set delivers the highest specificity and efficiency for the most reliable and accurate results. LNA enhancement enables Tm normalization across the panel giving the primers and probes higher binding affinity, dramatically increasing assay sensitivity and specificity.
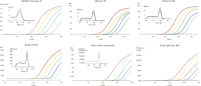
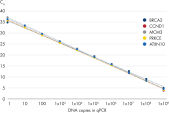
The increased sensitivity ensures excellent amplification efficiencies down to 1 RNA molecule, making it easier for you to detect low-abundance targets such as lncRNAs from less starting material (see figure QuantiNova LNA Probe PCR Assays provide accurate, sensitive and linear quantification of targets over a wide dynamic range and figure QuantiNova LNA Probe PCR Assays enable both high-expression and low-expression detection of mRNA). Increased specificity from the clever placement of LNA gives you a high signal-to-noise ratio, allowing discrimination of sequences that differ by only a single nucleotide and eliminating non-specific amplification and primer-dimer formation.
The high binding affinity of LNA bases increases the flexibility of primer and probe placement on the transcript, so we are able to use intelligent positioning to design assays for otherwise difficult-to-analyze targets. This gives you robust and reliable quantification, even for AU-rich targets, low-abundance transcripts, targets with high secondary structure and highly complex samples.
Principle
Robust and high-performance hydrolysis probe-based qPCR
QuantiNova LNA Probe PCR Custom Panels use FAM-labeled hydrolysis probe-based detection, which is highly sensitive and robust, because only the desired PCR product is detected. Probe-based detection still requires high PCR specificity to prevent amplification artifacts such as non-specific PCR products and primer–dimers that can compete for reaction components and compromise performance. However, we’ve already optimized the QuantiNova LNA Probe PCR system for you to eliminate non-specific amplification and give you accurate and sensitive detection every time.
Most comprehensive and specific coverage
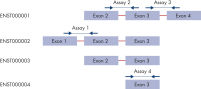
Our proprietary algorithm has been used to design over 1.3 million QuantiNova LNA Probe PCR Assays to provide the most sensitive, accurate and effective mRNA and lncRNA analysis. The predesigned assays cover most transcripts in the Ensembl database for human, mouse and rat genes, enabling PCR-based gene expression studies in the greatest depth possible. Most of the assays are intron-spanning when possible and detect only RNA. Assays that do not span an intron are designated as such, and if there is one exon in the target, unwanted signals can be easily eliminated using the QuantiNova Reverse Transcription Kit with the integrated gDNA removal step.
Custom Panel vs. Flexible Panel
QuantiNova LNA Probe PCR Custom Panels are a fully customizable product and are available in 96-well plate, 384-well plate and 100-well disc formats, with all of our different plate layouts available. The custom builder allows you to upload your own list of targets and add any predesigned or custom assays of your choice. Any reference genes and control assays and controls can also be added.
QuantiNova LNA Probe PCR Flexible Panels are a semi-customizable product and are available in 96-well plate, 384-well plate and 100-well disc formats, with up to 96 different assays on each plate. Flexible Panels can only contain assays from our available Focus Panels. Our convenient custom builder allows you to simply load the plate content of an existing Focus Panel and swap out the assays to meet your requirements. Reference gene assays and controls from our Focus Panels can also be added.
Available plate layouts, minimum order quantity and multiple plate discount
| Plate format | Number of different assays per plate | Configuration | Minimum order |
| 96-well or 100-well disc | 8 assays | 8 assays x 12 samples | 4 plates |
| 12 assays | 12 assays x 8 samples | 4 plates | |
| 16 assays | 16 assays x 6 samples | 4 plates | |
| 24 assays | 24 assays x 4 samples | 4 plates | |
| 32 assays | 32 assays x 3 samples | 4 plates | |
| 48 assays | 48 assays x 2 samples | 4 plates | |
| 96 assays | 96 assays x 1 sample | 6 plates | |
| 384-well | 8 assays | 8 assays x 48 samples | 6 plates |
| 12 assays | 12 assays x 32 samples | 6 plates | |
| 16 assays | 16 assays x 24 samples | 6 plates | |
| 24 assays | 24 assays x 16 samples | 6 plates | |
| 32 assays | 32 assays x 12 samples | 6 plates | |
| 48 assays (horizontal) | 48 assays x 8 samples | 6 plates | |
| 48 assays (normal vertical) | 48 assays x 8 samples | 6 plates | |
| 64 assays | 64 assays x 6 samples | 6 plates | |
| 96 assays | 96 assays x 4 samples | 6 plates | |
| 128 assays | 128 assays x 3 samples | 6 plates | |
| 192 assays | 192 assays x 2 samples | 12 plates | |
| 384 assays | 384 assays x 1 sample | 12 plates |
| Plate quantity ordered | Discount (%) |
| 1–5 | 0 |
| 6–11 | 15 |
| 12–23 | 30 |
| 24+ | 50 |
Reference Gene Assays for any study
A wide selection of human, mouse and rat Reference Gene Assays are available to enable high-quality data normalization and ensure reliable results. These assays target endogenous coding RNAs, long non-coding RNA and small nucleolar RNA molecules that are typically constitutively expressed in a wide variety of tissues. The Reference Gene Assays are FAM labeled and have been functionally validated as reference genes for the QuantiNova LNA PCR system and work optimally with the QuantiNova RT and PCR reagents.
Normalization of mRNA/lncRNA qPCR results
Normalization removes technical and biological inter-sample variation unrelated to the biological changes under investigation. Proper normalization is critical for correct analysis and interpretation of results from real-time PCR experiments. Most commonly, stably expressed reference genes are used for normalization.
It is generally recommended to test several endogenous control reference gene candidates before setting up your actual mRNA/lncRNA expression analysis. These candidates should be chosen from genes expected to be stably expressed over the whole range of samples under investigation. They could be stably expressed mRNAs or lncRNAs selected based on literature or preexisting data (e.g., NGS or qPCR panel screening). The QuantiNova LNA PCR system offers validated reference gene assays for RNAs that tend to be stably expressed and are therefore good candidates as reference genes.
All reference gene candidates should be empirically validated for each study. One option for normalizing PCR panel when profiling a large number of mRNAs/lncRNAs is to normalize against the global mean – the average of all expressed mRNAs/lncRNAs. This can be a good option in samples with a high call rate (expressed genes) but should be used with caution in samples with low call rates. It is also not a good option in samples for which the general gene expression level is changed. Further guidance on normalization can also be found in the GeneGlobe Data Analysis Center.
Complimentary data analysis
The complimentary, web-based data analysis tool in the GeneGlobe Data Analysis Center includes a user-friendly wizard to guide you step-by-step through the normalization and analysis of your data and generates publication-ready results figures.
Procedure
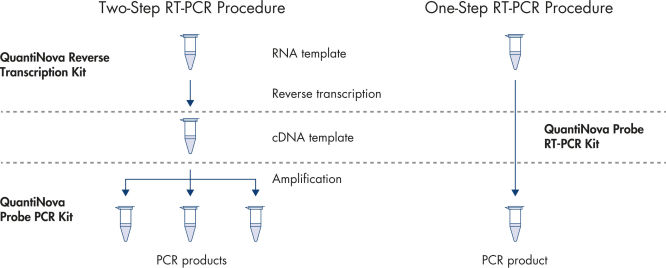
Choose one- or two-step qRT-PCR
QuantiNova LNA Probe PCR Custom Panels have been optimized using the time-tested and high-performance QuantiNova chemistry. Best results are obtained when reverse transcription is performed with the QuantiNova Reverse Transcription Kit. The resulting cDNA is then quantified by qPCR using the master mixes of the QuantiNova Probe PCR Kit combined with the assays on your panel.
For one-step qRT-PCR, we recommend using the QuantiNova Probe RT-PCR Kit, which offers quick reaction setup without need for optimization of reaction conditions. Simply combine RNA template with the ready-to-use RT-PCR master mix and the assays on your panel and start the reaction.
Fast and easy workflow
The fast and easy workflow can be completed in under 2 hours with minimal hands-on time and can be automated to save even more time and labor. The simple protocol has been tested for compatibility on all major brand qPCR cyclers (see figure QuantiNova LNA Probe PCR Assays deliver consistent performance across all thermal cycler instruments). Furthermore, the cDNA generated in the RT step can be used across the entire QuantiNova LNA Probe PCR System, allowing seamless transition from assays to panels depending on your research needs, saving you time and sample.
What you need to get started with the QuantiNova LNA Probe PCR system
Reverse transcription: QuantiNova Reverse Transcription Kit
qPCR mastermix: QuantiNova Probe PCR Kit
One-Step qRT-PCR (optional): QuantiNova Probe RT-PCR Kit
Assays or panels:
- QuantiNova LNA Probe PCR Assays
- QuantiNova LNA Probe PCR Reference Assays
- QuantiNova LNA Probe PCR Custom Assays
- QuantiNova LNA Probe PCR Focus Panels
- QuantiNova LNA Probe PCR lncRNA Focus Panels
- QuantiNova LNA Probe PCR Custom Panels
- QuantiNova LNA Probe PCR Flexible Panels
Applications
QuantiNova LNA Probe PCR Assays and Panels are highly suited for applications including:
- mRNA and lncRNA expression analysis, profiling and quantification
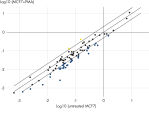
- Validation of RNA-seq gene expression data
- Gene expression profiling
- Signal and pathway analysis
- Confirming gene expression knockdown by LNA GapmeRs or siRNAs
- Biomarker development, including screening, identification and validation of disease-associated biomarkers
- Monitoring phenotypic changes related to gene expression
Supporting data and figures
Ultrafast one- or two-step procedure.