QIAquick Gel Extraction Kit
겔 또는 효소 반응에서 최대 10µg DNA(70bp~10kb)의 겔 추출/클린업에 사용합니다
겔 또는 효소 반응에서 최대 10µg DNA(70bp~10kb)의 겔 추출/클린업에 사용합니다
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Cat. No. / ID: 28704
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
QIAquick Gel Extraction Kit는 겔(최대 400mg 절편) 또는 효소 반응에서 DNA 절편의 실리카 막 기반 정제를 위한 스핀 컬럼, 완충액, collection 튜브를 제공합니다. 간단하고 빠른 결합-세척-용출 절차 및 30~50µl의 용출량으로 70bp~10kb 범위의 DNA를 정제합니다. 포함된 pH indicator 염료를 통해 스핀 컬럼에 결합하는 DNA에 대한 최적의 pH를 쉽게 파악할 수 있습니다. 또한 QIAquick PCR & Gel Cleanup Kit는 >100bp의 PCR 산물과 최대 10kb의 DNA를 정제하기 위한 완충액을 제공합니다. 해당 절차는 QIAcube Connect에서 완전히 자동화할 수 있습니다.
최적의 결과를 얻으려면 이 제품을 QIAvac 24 Plus와 함께 사용하는 것이 좋습니다.
QIAquick Gel Extraction Kit를 사용하면 샘플에서 뉴클레오타이드, 효소, 염, 아가로스, 브로민화 에티듐(ethidium bromide) 및 기타 불순물을 제거하여 최대 80%의 DNA를 회수할 수 있습니다(그림 " 겔에서 높은 회수율" 참조). 마이크로 원심분리기 또는 진공 매니폴드를 사용하여 1~24개 샘플에서 70bp~10kb 범위의 DNA를 정제합니다. 정제된 DNA는 예를 들어 염기서열 분석에 사용할 수 있습니다(그림 " 겔 추출 후 신뢰할 수 있는 염기서열 분석" 참조). 70bp보다 작거나 10kb보다 큰 DNA 절편은 QIAEX II Gel Extraction System으로 추출해야 합니다.
QIAquick PCR Purification 절차는 DNA 샘플에서 프라이머, 뉴클레오타이드, 효소, 미네랄 오일, 염 및 기타 불순물을 제거합니다(그림 " PCR 후 프라이머 완전 제거" 참조). 마이크로 원심분리기 또는 진공 매니폴드를 사용하여 100bp~10kb 범위의 DNA를 정제합니다.
QIAquick Kit에는 염도가 높은 완충액에서 DNA를 결합하고 저염 완충액 또는 물로 용출하기 위한 실리카 막 어셈블리가 포함되어 있습니다. 정제 절차는 DNA 샘플에서 프라이머, 뉴클레오타이드, 효소, 미네랄 오일, 염, 아가로스, 브로민화 에티듐(ethidium bromide) 및 기타 불순물을 제거합니다(그림 " 겔에서 높은 회수율" 참조). 실리카 막 기술은 분산 수지 및 슬러리(slurry)와 관련된 문제와 불편을 해소합니다. 특수 결합 완충액은 각 응용 분야에 최적화되어 있으며 특정 크기 범위 내에서 DNA 분자를 선택적으로 흡착하도록 합니다.
더 빠르고 편리한 샘플 처리 및 분석을 위해 겔 로딩 염료가 제공됩니다. GelPilot 로딩 염료는 아가로스 겔 실행 시간을 최적화하고 작은 DNA 절편이 너무 멀리 이동하는 것을 방지하기 위해 세 가지 추적 염료(크실렌 시아놀, 브로모페놀 블루, 오렌지 G)를 함유하고 있습니다(그림 "GelPilot 로딩 염료" 참조).
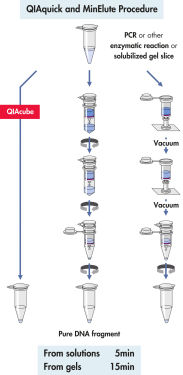
QIAquick system은 간단한 결합-세척-용출 절차를 사용합니다(순서도 " QIAquick 및 MinElute 절차" 참조). 겔 절편을 DNA 결합을 위한 최적의 pH를 쉽게 확인할 수 있는 pH indicator 염료가 포함된 완충액에서 용해하고, 혼합물을 QIAquick 스핀 컬럼에 적용합니다(그림 " pH indicator 염료" 참조). 핵산은 완충액의 높은 염도 조건에서 실리카 막에 흡착됩니다. 불순물을 씻어내고 제공된 소량의 저염 완충액 또는 물로 순수한 DNA를 용출하여 모든 후속 응용 분야에 바로 사용할 수 있습니다.
QIAquick 스핀 컬럼은 두 가지 편리한 취급 옵션을 제공하도록 고안되었습니다. 스핀 컬럼은 기존의 탁상용 마이크로 원심분리기 또는 루어 커넥터가 있는 모든 진공 매니폴드(예:QIAvac Luer Adapters가 있는 QIAvac 24 Plus)에 장착할 수 있습니다. QIAquick Gel Extraction Kit는 다른 QIAGEN 스핀 컬럼 기반 키트와 더불어 QIAcube Connect에서 완전히 자동화할 수 있어 생산성을 높이고 결과를 표준화할 수 있습니다(그림 "스핀 컬럼 취급 옵션 A, B, C, D" 및 " QIAcube Connect" 참조).
QIAquick system으로 정제된 DNA 절편은 염기서열 분석, 결찰(ligation) 및 형질전환, 제한효소 처리(restriction digestion), 라벨링, 미세 주입, PCR 및 체외 전사를 포함한 모든 응용 분야에서 바로 사용할 수 있습니다.
| Features | Specifications |
|---|---|
| Binding capacity | 10µg |
| Format | 튜브 |
| Fragment size | 70bp~10kb |
| Recovery: oligonucleotides dsDNA | 회수: dsDNA 절편 |
| Processing | 수동 |
| Removal <10mers 17–40mers dye terminator proteins | <10mers 제거 |
| Elution volume | 30~50µl |
| Technology | 실리카 기술 |
| Sample type: applications | DNA: PCR 반응 |