✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
REPLI-g Cell WGA & WTA Kit (12)
Cat no. / ID. 150052
✓ 24/7 automatic processing of online orders
✓ Knowledgeable and professional Product & Technical Support
✓ Fast and reliable (re)-ordering
Features
- Parallel amplification of genomic DNA and RNA (total or mRNA-enriched)
- Easily correlate genomic status with transcription pattern
- Negligible sequence bias due to MDA technology
- Amplify gDNA and RNA directly from cells or tissue
- For use with all downstream applications, including NGS
Product Details
Performance
Complete genomic and transcriptome coverage, with low experimental variability
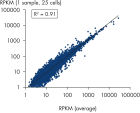
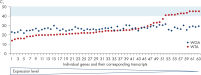
The REPLI-g Cell WGA & WTA Kit contains the novel REPLI-g SensiPhi DNA Polymerase, as well as an optimized set of buffers and reagents for parallel whole genome amplification (WGA) and whole transcriptome amplification (WTA) from 25–1000 cells, or equivalently small samples. Following efficient cell lysis, (and complete removal of genomic DNA followed by sensitive reverse transcription for the WTA reaction), the kit utilizes Multiple Displacement Amplification (MDA) technology for complete and unbiased amplification of gDNA and cDNA (see figure Multiple Displacement Amplification (MDA) technology). cDNA amplified using the REPLI-g Cell WGA & WTA Kit demonstrates a high degree of reproducibility from experiment to experiment, with minimal “technical noise”, ensuring highly reliable results (see figure High level of experimental reproducibility and minimized technical noise). The amplified gDNA and cDNA can be easily used in a variety of downstream applications (see figure REPLI-g amplified cDNA and gDNA perform like gDNA in downstream experiments).Ideal for analyzing the genomic and transcriptomic profiles of limited samples
The REPLI-g Cell WGA & WTA Kit provides highly uniform, parallel amplification across the entire genome and transcriptome, with negligible sequence bias. Even from samples as low as 25 cells, the kit ensures reliable whole genome and transcriptome amplification, making it highly suited for analyzing the direct consequences of genotypic changes on transcriptomes (see figure Highly suited for simultaneous genomic and transcriptomic analysis from the same limited sample).For use in a wide variety of research areas
The REPLI-g Cell WGA & WTA Kit efficiently amplifies gDNA and cDNA directly from cells and can be used with various samples that are often analyzed in clinical research, including tumor cells, stem cells, sorted cells, or other small samples (see table).| Sample material (cells/total RNA) | Research area |
|---|---|
| Cancer | Somatic genetic variant analysis |
| Tumor progression | |
| Tumor stem cells/evolution | |
| Tumor heterogeneity | |
| Somatic genetic variant analysis | |
| Human/animal | Biomarker research (SNPs, mutations, CNVs) |
| Mosaicism studies | |
| Stem cell research | |
| Genetic predisposition studies |
Principle
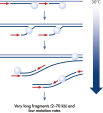
The REPLI-g Cell WGA & WTA Kit provides highly uniform amplification across the entire genome and transcriptome, with negligible sequence bias. The kit uses Multiple Displacement Amplification (MDA) technology, which carries out isothermal gDNA/cDNA amplification utilizing a uniquely processive DNA polymerase capable of replicating up to 70 kb without dissociating from the DNA template (see figure Multiple Displacement Amplification (MDA) technology).
Unique components of the REPLI-g Cell WGA & WTA Kit
- All of the kit’s enzymes and amplification components undergo a unique, controlled decontamination procedure to ensure elimination of REPLI-g amplifiable contaminating DNA or RNA. Following the procedure, the kits undergo stringent quality control to ensure complete functionality.
- The unique lysis buffer effectively stabilizes cellular RNA and DNA. This ensures that the resulting RNA accurately reflects the in vivo gene expression profile, and that RNA and DNA stays intact, maximizing genome coverage for subsequent analyses.
- All enzymatic steps have been specifically developed to enable efficient processing of RNA or DNA for accurate amplification. For example, these processes include effective gDNA removal prior to cDNA synthesis for RNA amplification and enzymatic DNA preparation that ensures efficient DNA amplification.
- Novel REPLI-g SensiPhi DNA Polymerase is used for Multiple Displacement Amplification (MDA). It is a newly developed, high-affinity enzyme that binds DNA more efficiently, especially when DNA concentration is low in the reaction mixture. In addition, and in contrast to PCR-based methods, REPLI-g SensiPhi DNA Polymerase has strong proofreading activity that results in 1000-fold fewer errors. It also has strong strand-displacement activity, enabling replication of DNA through stable hairpin structures that are resistant to Taq-based whole genome and whole transcriptome amplification procedures.
Procedure
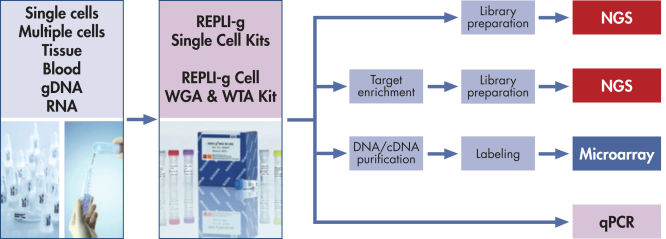
The REPLI-g Cell WGA & WTA Kit contains reagents for the following sequential reactions (see figure REPLI-g Cell WGA & WTA Kit procedure):
- Lysis of cells: a sample (containing 25–1000 cells) is lysed efficiently within 5 minutes, with no effect on RNA or DNA integrity. Lysed cells are divided into two aliquots of equal volume. The first aliquot is used for WGA, while the second aliquot is used for WTA of total RNA, or alternatively, mRNA-enriched (poly A+) RNA.
- Whole genome amplification: following cell lysis, DNA from one of the aliquots is modified for high efficiency ligation. Due to the nature of the ligation reaction, DNA fragments are not assembled in the order in which they would have originally existed in the organism. However, this does not affect detection of nucleic acid sequences (e.g., polymorphisms) in downstream applications, such as NGS, array analysis, or qPCR. After ligation, amplification using MDA technology and the novel, high-affinity REPLI-g SensiPhi DNA Polymerase takes place.
- Whole transcriptome amplification: following cell lysis, gDNA is removed from the second aliquot prior to starting the WTA process, since accurate measurement of transcript levels depends on the elimination of false-positive results caused by gDNA contamination. cDNA is prepared for high efficiency ligation and amplification. Depending on the primer chosen during the subsequent reverse transcription reaction, all transcripts for total RNA enrichment or only polyadenylated transcripts for poly A+ mRNA enrichment will be amplified.
Applications
The REPLI-g Cell WGA & WTA Kit allows uniform parallel amplification of all transcripts and genomic regions from one very small sample. gDNA and cDNA amplified in parallel by the kit are highly suited for next-generation sequencing (both DNA-Seq and RNA-Seq*), analysis using arrays, genotyping applications, or quantitative PCR.
The REPLI-g Cell WGA & WTA Kit is suitable for amplification of:
Transcriptome (RNA):
- mRNA with poly A+ tail
- Total RNA
- All regions of RNA transcripts
- 3’ ends of mRNAs
- Lnc and linc RNAs
Genome (DNA):
- Genomic DNA
- Mitochondrial DNA
It is not suitable for use with small nucleic acids (for example):
- tRNAs and miRNAs
- Strongly degraded DNA or RNA
Supporting data and figures
REPLI-g amplified cDNA and gDNA perform like gDNA in downstream experiments.